Target Information
| Target General Information | Top | |||||
|---|---|---|---|---|---|---|
| Target ID |
T91331
(Former ID: TTDC00131)
|
|||||
| Target Name |
Fibroblast growth factor receptor 3 (FGFR3)
|
|||||
| Synonyms |
JTK4; FGFR-3; CD333
Click to Show/Hide
|
|||||
| Gene Name |
FGFR3
|
|||||
| Target Type |
Successful target
|
[1] | ||||
| Disease | [+] 3 Target-related Diseases | + | ||||
| 1 | Liver cancer [ICD-11: 2C12] | |||||
| 2 | Coronary atherosclerosis [ICD-11: BA80] | |||||
| 3 | Myocardial infarction [ICD-11: BA41-BA43] | |||||
| Function |
Plays an essential role in the regulation of chondrocyte differentiation, proliferation and apoptosis, and is required for normal skeleton development. Regulates both osteogenesis and postnatal bone mineralization by osteoblasts. Promotes apoptosis in chondrocytes, but can also promote cancer cell proliferation. Required for normal development of the inner ear. Phosphorylates PLCG1, CBL and FRS2. Ligand binding leads to the activation of several signaling cascades. Activation of PLCG1 leads to the production of the cellular signaling molecules diacylglycerol and inositol 1,4,5-trisphosphate. Phosphorylation of FRS2 triggers recruitment of GRB2, GAB1, PIK3R1 and SOS1, and mediates activation of RAS, MAPK1/ERK2, MAPK3/ERK1 and the MAP kinase signaling pathway, as well as of the AKT1 signaling pathway. Plays a role in the regulation of vitamin D metabolism. Mutations that lead to constitutive kinase activation or impair normal FGFR3 maturation, internalization and degradation lead to aberrant signaling. Over-expressed or constitutively activated FGFR3 promotes activation of PTPN11/SHP2, STAT1, STAT5A and STAT5B. Secreted isoform 3 retains its capacity to bind FGF1 and FGF2 and hence may interfere with FGF signaling. Tyrosine-protein kinase that acts as cell-surface receptor for fibroblast growth factors and plays an essential role in the regulation of cell proliferation, differentiation and apoptosis.
Click to Show/Hide
|
|||||
| BioChemical Class |
Kinase
|
|||||
| UniProt ID | ||||||
| EC Number |
EC 2.7.10.1
|
|||||
| Sequence |
MGAPACALALCVAVAIVAGASSESLGTEQRVVGRAAEVPGPEPGQQEQLVFGSGDAVELS
CPPPGGGPMGPTVWVKDGTGLVPSERVLVGPQRLQVLNASHEDSGAYSCRQRLTQRVLCH FSVRVTDAPSSGDDEDGEDEAEDTGVDTGAPYWTRPERMDKKLLAVPAANTVRFRCPAAG NPTPSISWLKNGREFRGEHRIGGIKLRHQQWSLVMESVVPSDRGNYTCVVENKFGSIRQT YTLDVLERSPHRPILQAGLPANQTAVLGSDVEFHCKVYSDAQPHIQWLKHVEVNGSKVGP DGTPYVTVLKTAGANTTDKELEVLSLHNVTFEDAGEYTCLAGNSIGFSHHSAWLVVLPAE EELVEADEAGSVYAGILSYGVGFFLFILVVAAVTLCRLRSPPKKGLGSPTVHKISRFPLK RQVSLESNASMSSNTPLVRIARLSSGEGPTLANVSELELPADPKWELSRARLTLGKPLGE GCFGQVVMAEAIGIDKDRAAKPVTVAVKMLKDDATDKDLSDLVSEMEMMKMIGKHKNIIN LLGACTQGGPLYVLVEYAAKGNLREFLRARRPPGLDYSFDTCKPPEEQLTFKDLVSCAYQ VARGMEYLASQKCIHRDLAARNVLVTEDNVMKIADFGLARDVHNLDYYKKTTNGRLPVKW MAPEALFDRVYTHQSDVWSFGVLLWEIFTLGGSPYPGIPVEELFKLLKEGHRMDKPANCT HDLYMIMRECWHAAPSQRPTFKQLVEDLDRVLTVTSTDEYLDLSAPFEQYSPGGQDTPSS SSSGDDSVFAHDLLPPAPPSSGGSRT Click to Show/Hide
|
|||||
| 3D Structure | Click to Show 3D Structure of This Target | AlphaFold | ||||
| Drugs and Modes of Action | Top | |||||
|---|---|---|---|---|---|---|
| Approved Drug(s) | [+] 2 Approved Drugs | + | ||||
| 1 | Pemigatinib | Drug Info | Approved | Cholangiocarcinoma | [2] | |
| 2 | Trapidil | Drug Info | Phase 4 | Coronary artery disease | [3] | |
| Clinical Trial Drug(s) | [+] 10 Clinical Trial Drugs | + | ||||
| 1 | BMS-582664 | Drug Info | Phase 3 | Hepatocellular carcinoma | [4], [5] | |
| 2 | E-3810 | Drug Info | Phase 3 | Solid tumour/cancer | [6], [7] | |
| 3 | TKI258 | Drug Info | Phase 3 | Renal cell carcinoma | [8] | |
| 4 | B-701 | Drug Info | Phase 2 | Bladder cancer | [9], [10] | |
| 5 | Debio 1347 | Drug Info | Phase 2 | Solid tumour/cancer | [11] | |
| 6 | Recifercept | Drug Info | Phase 2 | Achondroplasia | [12] | |
| 7 | AEE-788 | Drug Info | Phase 1/2 | Solid tumour/cancer | [13], [14] | |
| 8 | MK-2461 | Drug Info | Phase 1/2 | Alzheimer disease | [15] | |
| 9 | Anti-FGFR3 | Drug Info | Phase 1 | Multiple myeloma | [16] | |
| 10 | SAR442501 | Drug Info | Phase 1 | Achondroplasia | [17] | |
| Discontinued Drug(s) | [+] 1 Discontinued Drugs | + | ||||
| 1 | PD-0183812 | Drug Info | Terminated | Retinoblastoma | [18] | |
| Mode of Action | [+] 2 Modes of Action | + | ||||
| Inhibitor | [+] 14 Inhibitor drugs | + | ||||
| 1 | Pemigatinib | Drug Info | [2] | |||
| 2 | Trapidil | Drug Info | [1] | |||
| 3 | BMS-582664 | Drug Info | [19] | |||
| 4 | E-3810 | Drug Info | [20] | |||
| 5 | TKI258 | Drug Info | [19] | |||
| 6 | Debio 1347 | Drug Info | [9] | |||
| 7 | AEE-788 | Drug Info | [19] | |||
| 8 | MK-2461 | Drug Info | [15] | |||
| 9 | PD-0183812 | Drug Info | [22] | |||
| 10 | 5,11-Dimethyl-6H-pyrido[4,3-b]carbazol-9-ol | Drug Info | [23] | |||
| 11 | ACTB-1003 | Drug Info | [24] | |||
| 12 | PMID21493067C1d | Drug Info | [25] | |||
| 13 | Ro-4396686 | Drug Info | [26] | |||
| 14 | SU5402 | Drug Info | [1] | |||
| Antagonist | [+] 1 Antagonist drugs | + | ||||
| 1 | B-701 | Drug Info | [10] | |||
| Cell-based Target Expression Variations | Top | |||||
|---|---|---|---|---|---|---|
| Cell-based Target Expression Variations | ||||||
| Drug Binding Sites of Target | Top | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Ligand Name: L-betagamma-meATP | Ligand Info | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Structure Description | Crystal Structure of an Asymmetric Dimer of FGF Receptor 3 Kinases Trapped in A-loop Tyrosine Transphosphorylation Reaction | PDB:6PNX | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Method | X-ray diffraction | Resolution | 2.20 Å | Mutation | Yes | [27] | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PDB Sequence |
ELELPADPKW
465 ELSRARLTLG475 KPLGEGAFGQ485 VVMAEAIGID495 KDRAAKPVTV505 AVKMLKDDAT 515 DKDLSDLVSE525 MEMMKMIGKH535 KNIINLLGAC545 TQGGPLYVLV555 EYAAKGNLRE 565 FLRARRPPPE586 EQLTFKDLVS596 CAYQVARGME606 YLASQKCIHR616 DLAARNVLVT 626 EDNVMKIADF636 GLARDVHNLD646 YYKKTTNGRL656 PVKWMAPEAL666 FDEVYTHQSD 676 VWSFGVLLWE686 IFTLGGSPYP696 GIPVEELFKL706 LKEGHRMDKP716 ANCTHDLYMI 726 MRECWHAAPS736 QRPTFKQLVE746 DLDRVLTVTS756 T
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
☰6PNX Nodes: OProtein ▢Nucleotide ◇Chemical ▢Biopolymer Lines: Interactions at 4 Å Dynamically generated for selected residues. Nodes can be dragged or clicked. Label: Selection: Name:
Note: VAST+ finds other macromolecular structures that have a similar biological unit. To do this, VAST+ takes into consideration the complete set of 3D domains that VAST identified within a query structure, throughout all of its component protein molecules, and finds other macromolecular structures that have a similar set of proteins/3D domains. PDB ID: Note: VAST identifies 3D domains (substructures) within each protein structure in the Molecular Modeling Database (MMDB), and then finds other protein structures that have one or more similar 3D domains, using purely geometric criteria. You have two ways to do a VAST search. Option 1, search with your selection (all residues are selected by default) in the loaded structures: Option 2, search with PDB ID and chain name: PDB ID: Chain Name: Option 3, search with a PDB file: 1. your selection (all residues are selected by default) in the loaded structures to Foldseek web server. 2 (Optional). Once you see the structure neighbors, you can view the alignment in iCn3D by inputing a list of PDB chain IDs or AlphaFold UniProt IDs below. The PDB chain IDs are the same as the record names such as "1HHO_A". The UniProt ID is the text between "AF-" and "-F1". For example, the UniProt ID for the record name "AF-P69905-F1-model_v4" is "P69905". Chain ID List: BCIF/MMTF ID: PDB ID: Note: AlphaFold produces a per-residue confidence score (pLDDT) between 0 and 100: Very high (pLDDT > 90) Confident (90 > pLDDT > 70) Low (70 > pLDDT > 50) Very low (pLDDT < 50) AlphaFold Uniprot ID: PAE Map: NCBI Protein Accession: Note: Several PDB files could be concatenated into a single PDB file. Use the line "ENDMDL" to separate PDB files. PDB File: Multiple PDB Files: The custom JSON file on residue colors has the following format for proteins("ALA" and "ARG") and nucleotides("G" and "A"): {"ALA":"#C8C8C8", "ARG":"#145AFF", ..., "G":"#008000", "A":"#6080FF", ...} Residue Color File: The custom file for the structure has two columns separated by space or tab: residue number, and score in the range of 0-100. If you click "Apply Custom Color" button, the scores 0, 50 and 100 correspond to the three colors specified below. If you click "Apply Custom Tube", the selected residues will be displayed in a style similar to "B-factor Tube". Custom File: 1. Score to Color: 0: 50: 100: or 2. You can define your own reference numbers in a custom file using Excel, and then export it as a CSV file. An example file is shown below with cells separated by commas. refnum,11,12,,21,22,,10C,11C,20CThe first row defines the reference residue numbers, which could be any strings. The 1st cell could be anything. The rest cells are reference residue numbers (e.g., 11, 21, 10C, etc.) or empty cells. Each chain has a separate row. The first cell of the second row is the chain ID "1TUP_A". The rest cells are the corresponding real residue numbers for reference residue numbers in the first row. For example, the reference numbers for residues 100, 101, and 132 in the chain 1TUP_A are 11, 12, and 22, respectively. The fourth row shows another set of reference numners for the chain "1TUP_C". It could be a chain from a different structure. To select all residues corresponding to the reference numbers, you can simplay replace ":" with "%" in the Specification. For example, "%12" selects the residue 101 in 1TUP_A and the residue 111 in 1TUP_B. ".A%12" has the chain "A" filter and selects the residue 101 in 1TUP_A. Custom File: Enter the PDB IDs or MMDB IDs of the structures: ID1: ID2: VAST+ based on VAST: VAST+ based on TM-align: All chains will be aligned to the first chain in the comma-separated chain IDs. Each chain ID has the form of PDBID_chain (e.g., 1HHO_A, case sensitive) or UniprotID (e.g., P69905 for AlphaFold structures). Chain IDs: (Note: To align chains in custom PDB files, you could load them in "File > Open File > PDB Files (appendable)" and click "Analysis > Defined Sets". Finally select multiple chains in Defined Sets and click "File > Realign Selection".) All chains will be aligned to the first chain in the comma-separated chain IDs. Each chain ID has the form of PDBID_chain (e.g., 1HHO_A, case sensitive) or UniprotID (e.g., P69905 for AlphaFold structures). Chain IDs: The sequence alignment (followed by structure alignment) is based on residue numbers in the First/Master chain: (Note: To align chains in custom PDB files, you could load them in "File > Open File > PDB Files (appendable)" and click "Analysis > Defined Sets". Finally select multiple chains in Defined Sets and click "File > Realign Selection".) All chains will be aligned to the first chain in the comma-separated chain IDs. Each chain ID has the form of PDBID_chain (e.g., 1HHO_A, case sensitive) or UniprotID (e.g., P69905 for AlphaFold structures). Chain IDs: Each alignment is defined as " | "-separated residue lists in one line. "10-50" means a range of residues from 10 to 50. Option 1: Option 2: All chains will be aligned to the first chain in the comma-separated chain IDs. Each chain ID has the form of PDBID_chain (e.g., 1HHO_A, case sensitive) or UniprotID (e.g., P69905 for AlphaFold structures). Chain IDs: Each alignment is defined as " | "-separated residue lists in one line. "10-50" means a range of residues from 10 to 50. Please specify the mutations with a comma separated mutation list. Each mutation can be specified as "[uppercase PDB ID or AlphaFold UniProt ID]_[Chain Name]_[Residue Number]_[One Letter Mutant Residue]". E.g., the mutation of N501Y in the E chain of PDB 6M0J can be specified as "6M0J_E_501_Y". For AlphaFold structures, the "Chain ID" is "A". If you load a custom structure without PDB or UniProt ID, you can open "Seq. & Annotations" window and find the chain ID such as "stru_A". The part before the underscore is the structure ID, which can be used to specify the mutation such as "stru_A_...". Remember to choose "Show Mutation in: Current Page". Mutations: ID Type: PDB IDAlphaFold UniProt ID Show Mutation in: Current PageNew Page Mol2 File: SDF File: XYZ File: AlphaFold PAE File: File type: URL in the same host: Multiple mmCIF Files: mmCIF ID: MMDB or PDB ID: Note: The "biological unit" is the biochemically active form of a biomolecule, List of PDB, MMDB, or AlphaFold UniProt structures: or Note: The "biological unit" is the biochemically active form of a biomolecule, Enter a protein sequence ID (or FASTA sequence) and the aligned protein accession, which can be found using the BLAST search with the protein sequence ID or FASTA sequence as input. If the protein accession is not a PDB chain, the corresponding AlphaFold UniProt structure is used. Protein Sequence ID(NCBI protein accession of a sequence): or FASTA sequence: Aligned Protein Accession (or a chain of a PDB): The sequence to structure prediction is done via ESM Metagenomic Atlas. The sequence should be less than 400 characters. For any sequence longer than 400, please see the discussion here. FASTA sequence: Your note will be saved in the HTML file when you click "File > Save File > iCn3D PNG Image". Protein/Gene name: PubChem CID/Name/InchI: Chemical SMILES: Multiple iCn3D PNG images: State file: Since January 6, 2021, you can show the original view with the archived version of iCn3D by pasting your URL below and click "Show Originial View". Note the version in the parameter "v" was used to replace "full.html" with "full_[v].html" in the URL. Share Link URL: Selection file: Collection File: Structures: Note: Always load a PDB file before loading map files. If you don't specify the threshold below, a default one will be chosen. 2fofc contour at default threshold or at: σ fofc contour at default threshold or at: σ Note: Always load a PDB file before loading map files. If you don't specify the threshold below, a default one will be chosen. 2fofc contour at default threshold or at: σ URL in the same host: fofc contour at default threshold or at: σ URL in the same host: Click in the input box to use the color picker: Custom Color: Grid Size: Salt Concentration: M Potential contour at: kT/e(25.6mV at 298K) Note: Only the selected residues are used for DelPhi potential calculation by solving linear Poisson-Boltzmann equation. Grid Size: Salt Concentration: M Surface with max potential at: kT/e(25.6mV at 298K) Surface: Opacity: Wireframe: Note: Only the selected residues are used for DelPhi potential calculation by solving linear Poisson-Boltzmann equation. Potential contour at: kT/e(25.6mV at 298K) Note: Always load a PDB file before loading a PQR or DelPhi potential file. Potential contour at: kT/e(25.6mV at 298K) Grid Size: Salt Concentration: M PQR URL in the same host: Phi URL in the same host: Cube URL in the same host: Note: Always load a PDB file before loading a PQR or DelPhi potential file. Symmetry: Distance: Contact Type: 1. Choose interaction types and their thresholds:
4. Sort Interactions on: to show two lines of residue nodes to show map with atom details to show interactions with strength parameters in 0-200:
(Note: you can also adjust thresholds at #1 to add/remove interactions.) 5. and select new sets 1. Select sets below or use your current selection: 2. 1. Select sets below or use your current selection. 2. 1. Select sets below or use your current selection: 2. Overall maximum RMSD: Å 3. 1. Select sets below: 2. 1. Select sets below: 2. 1. Select sets below: 2. 1. Select sets below: 2. Hold Ctrl key to select multiple nodes/lines. Green: H-Bonds; Cyan: Salt Bridge/Ionic; Grey: Contacts Magenta: Halogen Bonds; Red: π-Cation; Blue: π-Stacking Scale: Hold Ctrl key to select multiple nodes. Scale: Note: Nodes/Residues can be dragged. Both nodes and dashed lines/interactions can be clicked to select residues. Color legend for interactions (dashed lines): Green: H-Bonds; Cyan: Salt Bridge/Ionic; Grey: Contacts Magenta: Halogen Bonds; Red: π-Cation; Blue: π-Stacking Scale: Hold Ctrl key to select multiple nodes. Scale: Hold Ctrl key to select multiple nodes. Scale:
Contour at: σ Contour at: σ Contour at: % of maximum EM values 1. Select the first set: 2. Sphere with a radius: Å 3. Select the second set to apply the sphere: 4. the sphere around the first set of atoms interacting/contacting residue pairs in a file Note: The membranes are parallel to the X-Y plane. The center of the membranes is at Z = 0. 1. Extracellular membrane Z-axis position: Å 2. intracellular membrane Z-axis position: Å 3. the adjusted membranes Note: The membranes are parallel to the X-Y plane. The center of the membranes is at Z = 0. 1. Z-axis position of the first X-Y plane: Å 2. Z-axis position of the second X-Y plane: Å 3. the region between the planes to Defined Sets 1. Text: 2. Size: 3. Color: 4. Pick TWO atoms while holding "Alt" key 5. 1. Text: 2. Size: 3. Color: 4. Color for all labels: 1. Pick TWO atoms while holding "Alt" key 2. Line Color: 3. 1. Pick TWO atoms while holding "Alt" key 2. Color: 3. 1. Select two sets
3. 1. Select two sets
2. Line style: 3. Line radius: 4. Color: 5. Opacity: 6. 1. Select a set: 2. Shape: 3. Radius: 4. Color: 5. Opacity: 6. 1. Select sets for pairwise distances
Note: Each set is represented by a vector, which is the X-axis of the principle axes. The angles between the vectors are then calculated. 1. Select sets for pairwise angles
1. Pick TWO atoms while holding "Alt" key 2. Line Radius: (for stabilizers, hydrogen bonds, distance lines, default 0.1) Coil Radius: (for coils, default 0.3) Stick Radius: (for sticks, default 0.4) Cross-Linkage Radius: (for cross-linkages, default 0.4) Trace Radius: (for C alpha trace, O3' trace, default 0.4) Ribbon Thickness: (for helix and sheet ribbons, nucleotide ribbons, default 0.2) Protein Ribbon Width: (for helix and sheet ribbons, default 1.3) Nucleotide Ribbon Width: (for nucleotide ribbons, default 0.8) Ball Scale: (for styles 'Ball and Stick' and 'Dot', default 0.3) Note: The following parameters will be saved in cache. You just need to set them once. 1. Shininess: (for the shininess of the 3D objects, default 40) 2. Three directional lights: Key Light: (for the light strength of the key light, default 0.8) Fill Light: (for the light strength of the fill light, default 0.4) Back Light: (for the light strength of the back light, default 0.2) 3. Thickness: Line Radius: (for stabilizers, hydrogen bonds, distance lines, default 0.1) Coil Radius: (for coils, default 0.3) Stick Radius: (for sticks, default 0.4) Cross-Linkage Radius: (for cross-linkages, default 0.4) Trace Radius: (for C alpha trace, O3' trace, default 0.4) Ribbon Thickness: (for helix and sheet ribbons, nucleotide ribbons, default 0.2) Protein Ribbon Width: (for helix and sheet ribbons, default 1.3) Nucleotide Ribbon Width: (for nucleotide ribbons, default 0.8) Ball Scale: (for styles 'Ball and Stick' and 'Dot', default 0.3) 4. Show Glycan Cartoon: (0: hide, 1: show, default 0) 5. Show Membrane: (0: hide, 1: show, default 1) 6. Enlarge Command Window: (0: Regular, 1: Large, default 0) Name: 1. URLs Used in Browsers Please copy one of the URLs below. They show the same result. (To add a title to share link, click "Windows > Your Note" and click "File > Share Link" again.) Original URL with commands: Lifelong Short URL:(To replace this URL, send a pull request to update share.html at iCn3D GitHub) Lifelong Short URL + Window Title:(To update the window title, click "Analysis > Your Note/Window Title".) 2. Commands Used in Jupyter Noteboook Please copy the following commands into a cell in Jupyter Notebook to show the same result. More details are at https://github.com/ncbi/icn3d/tree/master/jupyternotebook. Annotations:
Zoom: mouse wheel; Move: left button; Select Multiple Nodes: Ctrl Key and drag an Area Force on Nodes: Label Size: Internal Edges: Solvent Accessible Surface Area(SASA) calculated using the EDTSurf algorithm: (0-20% out is considered "in". 50-100% out is considered "out".) Toal: Å2 Color each residue based on the percentage of solvent accessilbe surface area. The color ranges from blue, to white, to red for a percentage of 0, 35(variable), and 100, respectively. Middle Percentage(White): % Select residue based on the percentage of solvent accessilbe surface area. The values are in the range of 0-100. Min Percentage: % Max Percentage: % Select residue based on B-factor/pLDDT. The values are in the range of 0-100. Min B-factor/pLDDT: % Max B-factor/pLDDT: % X: Y: Z: Vector 1, X: Y: Z: Vector 2, X: Y: Z: The angle is: degree. 0: 4: 8: 12: 1: 5: 9: 13: 2: 6: 10: 14: 3: 7: 11: 15: Choose an Ig template for selected residues: Choose an Ig template to align with selected residues: |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Click to View More Binding Site Information of This Target and Ligand Pair | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Ligand Name: Phosphonotyrosine | Ligand Info | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Structure Description | Crystal structure of FGFR3 in complex with pyrimidine derivative | PDB:6LVM | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Method | X-ray diffraction | Resolution | 2.53 Å | Mutation | No | [28] | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PDB Sequence |
PEDPKWEFPR
469 DKLTLGKPLG479 EGCFGQVVMA489 EAIGIDKDRA499 AKPVTVAVKM509 LKDDATDKDL 519 SDLVSEMEMM529 KMIGKHKNII539 NLLGACTQGG549 PLYVLVEYAA559 KGNLREFLRA 569 RRPPGLDSEQ588 LTFKDLVSCA598 YQVARGMEYL608 ASQKCIHRDL618 AARNVLVTED 628 NVMKIADFGL638 ARDVHNLDKK650 TTNGRLPVKW660 MAPEALFDRV670 YTHQSDVWSF 680 GVLLWEIFTL690 GGSPYPGIPV700 EELFKLLKEG710 HRMDKPANCT720 HDLYMIMREC 730 WHAAPSQRPT740 FKQLVEDLDR750 VLTVT
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
☰6LVM Nodes: OProtein ▢Nucleotide ◇Chemical ▢Biopolymer Lines: Interactions at 4 Å Dynamically generated for selected residues. Nodes can be dragged or clicked. Label: Selection: Name:
Note: VAST+ finds other macromolecular structures that have a similar biological unit. To do this, VAST+ takes into consideration the complete set of 3D domains that VAST identified within a query structure, throughout all of its component protein molecules, and finds other macromolecular structures that have a similar set of proteins/3D domains. PDB ID: Note: VAST identifies 3D domains (substructures) within each protein structure in the Molecular Modeling Database (MMDB), and then finds other protein structures that have one or more similar 3D domains, using purely geometric criteria. You have two ways to do a VAST search. Option 1, search with your selection (all residues are selected by default) in the loaded structures: Option 2, search with PDB ID and chain name: PDB ID: Chain Name: Option 3, search with a PDB file: 1. your selection (all residues are selected by default) in the loaded structures to Foldseek web server. 2 (Optional). Once you see the structure neighbors, you can view the alignment in iCn3D by inputing a list of PDB chain IDs or AlphaFold UniProt IDs below. The PDB chain IDs are the same as the record names such as "1HHO_A". The UniProt ID is the text between "AF-" and "-F1". For example, the UniProt ID for the record name "AF-P69905-F1-model_v4" is "P69905". Chain ID List: BCIF/MMTF ID: PDB ID: Note: AlphaFold produces a per-residue confidence score (pLDDT) between 0 and 100: Very high (pLDDT > 90) Confident (90 > pLDDT > 70) Low (70 > pLDDT > 50) Very low (pLDDT < 50) AlphaFold Uniprot ID: PAE Map: NCBI Protein Accession: Note: Several PDB files could be concatenated into a single PDB file. Use the line "ENDMDL" to separate PDB files. PDB File: Multiple PDB Files: The custom JSON file on residue colors has the following format for proteins("ALA" and "ARG") and nucleotides("G" and "A"): {"ALA":"#C8C8C8", "ARG":"#145AFF", ..., "G":"#008000", "A":"#6080FF", ...} Residue Color File: The custom file for the structure has two columns separated by space or tab: residue number, and score in the range of 0-100. If you click "Apply Custom Color" button, the scores 0, 50 and 100 correspond to the three colors specified below. If you click "Apply Custom Tube", the selected residues will be displayed in a style similar to "B-factor Tube". Custom File: 1. Score to Color: 0: 50: 100: or 2. You can define your own reference numbers in a custom file using Excel, and then export it as a CSV file. An example file is shown below with cells separated by commas. refnum,11,12,,21,22,,10C,11C,20CThe first row defines the reference residue numbers, which could be any strings. The 1st cell could be anything. The rest cells are reference residue numbers (e.g., 11, 21, 10C, etc.) or empty cells. Each chain has a separate row. The first cell of the second row is the chain ID "1TUP_A". The rest cells are the corresponding real residue numbers for reference residue numbers in the first row. For example, the reference numbers for residues 100, 101, and 132 in the chain 1TUP_A are 11, 12, and 22, respectively. The fourth row shows another set of reference numners for the chain "1TUP_C". It could be a chain from a different structure. To select all residues corresponding to the reference numbers, you can simplay replace ":" with "%" in the Specification. For example, "%12" selects the residue 101 in 1TUP_A and the residue 111 in 1TUP_B. ".A%12" has the chain "A" filter and selects the residue 101 in 1TUP_A. Custom File: Enter the PDB IDs or MMDB IDs of the structures: ID1: ID2: VAST+ based on VAST: VAST+ based on TM-align: All chains will be aligned to the first chain in the comma-separated chain IDs. Each chain ID has the form of PDBID_chain (e.g., 1HHO_A, case sensitive) or UniprotID (e.g., P69905 for AlphaFold structures). Chain IDs: (Note: To align chains in custom PDB files, you could load them in "File > Open File > PDB Files (appendable)" and click "Analysis > Defined Sets". Finally select multiple chains in Defined Sets and click "File > Realign Selection".) All chains will be aligned to the first chain in the comma-separated chain IDs. Each chain ID has the form of PDBID_chain (e.g., 1HHO_A, case sensitive) or UniprotID (e.g., P69905 for AlphaFold structures). Chain IDs: The sequence alignment (followed by structure alignment) is based on residue numbers in the First/Master chain: (Note: To align chains in custom PDB files, you could load them in "File > Open File > PDB Files (appendable)" and click "Analysis > Defined Sets". Finally select multiple chains in Defined Sets and click "File > Realign Selection".) All chains will be aligned to the first chain in the comma-separated chain IDs. Each chain ID has the form of PDBID_chain (e.g., 1HHO_A, case sensitive) or UniprotID (e.g., P69905 for AlphaFold structures). Chain IDs: Each alignment is defined as " | "-separated residue lists in one line. "10-50" means a range of residues from 10 to 50. Option 1: Option 2: All chains will be aligned to the first chain in the comma-separated chain IDs. Each chain ID has the form of PDBID_chain (e.g., 1HHO_A, case sensitive) or UniprotID (e.g., P69905 for AlphaFold structures). Chain IDs: Each alignment is defined as " | "-separated residue lists in one line. "10-50" means a range of residues from 10 to 50. Please specify the mutations with a comma separated mutation list. Each mutation can be specified as "[uppercase PDB ID or AlphaFold UniProt ID]_[Chain Name]_[Residue Number]_[One Letter Mutant Residue]". E.g., the mutation of N501Y in the E chain of PDB 6M0J can be specified as "6M0J_E_501_Y". For AlphaFold structures, the "Chain ID" is "A". If you load a custom structure without PDB or UniProt ID, you can open "Seq. & Annotations" window and find the chain ID such as "stru_A". The part before the underscore is the structure ID, which can be used to specify the mutation such as "stru_A_...". Remember to choose "Show Mutation in: Current Page". Mutations: ID Type: PDB IDAlphaFold UniProt ID Show Mutation in: Current PageNew Page Mol2 File: SDF File: XYZ File: AlphaFold PAE File: File type: URL in the same host: Multiple mmCIF Files: mmCIF ID: MMDB or PDB ID: Note: The "biological unit" is the biochemically active form of a biomolecule, List of PDB, MMDB, or AlphaFold UniProt structures: or Note: The "biological unit" is the biochemically active form of a biomolecule, Enter a protein sequence ID (or FASTA sequence) and the aligned protein accession, which can be found using the BLAST search with the protein sequence ID or FASTA sequence as input. If the protein accession is not a PDB chain, the corresponding AlphaFold UniProt structure is used. Protein Sequence ID(NCBI protein accession of a sequence): or FASTA sequence: Aligned Protein Accession (or a chain of a PDB): The sequence to structure prediction is done via ESM Metagenomic Atlas. The sequence should be less than 400 characters. For any sequence longer than 400, please see the discussion here. FASTA sequence: Your note will be saved in the HTML file when you click "File > Save File > iCn3D PNG Image". Protein/Gene name: PubChem CID/Name/InchI: Chemical SMILES: Multiple iCn3D PNG images: State file: Since January 6, 2021, you can show the original view with the archived version of iCn3D by pasting your URL below and click "Show Originial View". Note the version in the parameter "v" was used to replace "full.html" with "full_[v].html" in the URL. Share Link URL: Selection file: Collection File: Structures: Note: Always load a PDB file before loading map files. If you don't specify the threshold below, a default one will be chosen. 2fofc contour at default threshold or at: σ fofc contour at default threshold or at: σ Note: Always load a PDB file before loading map files. If you don't specify the threshold below, a default one will be chosen. 2fofc contour at default threshold or at: σ URL in the same host: fofc contour at default threshold or at: σ URL in the same host: Click in the input box to use the color picker: Custom Color: Grid Size: Salt Concentration: M Potential contour at: kT/e(25.6mV at 298K) Note: Only the selected residues are used for DelPhi potential calculation by solving linear Poisson-Boltzmann equation. Grid Size: Salt Concentration: M Surface with max potential at: kT/e(25.6mV at 298K) Surface: Opacity: Wireframe: Note: Only the selected residues are used for DelPhi potential calculation by solving linear Poisson-Boltzmann equation. Potential contour at: kT/e(25.6mV at 298K) Note: Always load a PDB file before loading a PQR or DelPhi potential file. Potential contour at: kT/e(25.6mV at 298K) Grid Size: Salt Concentration: M PQR URL in the same host: Phi URL in the same host: Cube URL in the same host: Note: Always load a PDB file before loading a PQR or DelPhi potential file. Symmetry: Distance: Contact Type: 1. Choose interaction types and their thresholds:
4. Sort Interactions on: to show two lines of residue nodes to show map with atom details to show interactions with strength parameters in 0-200:
(Note: you can also adjust thresholds at #1 to add/remove interactions.) 5. and select new sets 1. Select sets below or use your current selection: 2. 1. Select sets below or use your current selection. 2. 1. Select sets below or use your current selection: 2. Overall maximum RMSD: Å 3. 1. Select sets below: 2. 1. Select sets below: 2. 1. Select sets below: 2. 1. Select sets below: 2. Hold Ctrl key to select multiple nodes/lines. Green: H-Bonds; Cyan: Salt Bridge/Ionic; Grey: Contacts Magenta: Halogen Bonds; Red: π-Cation; Blue: π-Stacking Scale: Hold Ctrl key to select multiple nodes. Scale: Note: Nodes/Residues can be dragged. Both nodes and dashed lines/interactions can be clicked to select residues. Color legend for interactions (dashed lines): Green: H-Bonds; Cyan: Salt Bridge/Ionic; Grey: Contacts Magenta: Halogen Bonds; Red: π-Cation; Blue: π-Stacking Scale: Hold Ctrl key to select multiple nodes. Scale: Hold Ctrl key to select multiple nodes. Scale:
Contour at: σ Contour at: σ Contour at: % of maximum EM values 1. Select the first set: 2. Sphere with a radius: Å 3. Select the second set to apply the sphere: 4. the sphere around the first set of atoms interacting/contacting residue pairs in a file Note: The membranes are parallel to the X-Y plane. The center of the membranes is at Z = 0. 1. Extracellular membrane Z-axis position: Å 2. intracellular membrane Z-axis position: Å 3. the adjusted membranes Note: The membranes are parallel to the X-Y plane. The center of the membranes is at Z = 0. 1. Z-axis position of the first X-Y plane: Å 2. Z-axis position of the second X-Y plane: Å 3. the region between the planes to Defined Sets 1. Text: 2. Size: 3. Color: 4. Pick TWO atoms while holding "Alt" key 5. 1. Text: 2. Size: 3. Color: 4. Color for all labels: 1. Pick TWO atoms while holding "Alt" key 2. Line Color: 3. 1. Pick TWO atoms while holding "Alt" key 2. Color: 3. 1. Select two sets
3. 1. Select two sets
2. Line style: 3. Line radius: 4. Color: 5. Opacity: 6. 1. Select a set: 2. Shape: 3. Radius: 4. Color: 5. Opacity: 6. 1. Select sets for pairwise distances
Note: Each set is represented by a vector, which is the X-axis of the principle axes. The angles between the vectors are then calculated. 1. Select sets for pairwise angles
1. Pick TWO atoms while holding "Alt" key 2. Line Radius: (for stabilizers, hydrogen bonds, distance lines, default 0.1) Coil Radius: (for coils, default 0.3) Stick Radius: (for sticks, default 0.4) Cross-Linkage Radius: (for cross-linkages, default 0.4) Trace Radius: (for C alpha trace, O3' trace, default 0.4) Ribbon Thickness: (for helix and sheet ribbons, nucleotide ribbons, default 0.2) Protein Ribbon Width: (for helix and sheet ribbons, default 1.3) Nucleotide Ribbon Width: (for nucleotide ribbons, default 0.8) Ball Scale: (for styles 'Ball and Stick' and 'Dot', default 0.3) Note: The following parameters will be saved in cache. You just need to set them once. 1. Shininess: (for the shininess of the 3D objects, default 40) 2. Three directional lights: Key Light: (for the light strength of the key light, default 0.8) Fill Light: (for the light strength of the fill light, default 0.4) Back Light: (for the light strength of the back light, default 0.2) 3. Thickness: Line Radius: (for stabilizers, hydrogen bonds, distance lines, default 0.1) Coil Radius: (for coils, default 0.3) Stick Radius: (for sticks, default 0.4) Cross-Linkage Radius: (for cross-linkages, default 0.4) Trace Radius: (for C alpha trace, O3' trace, default 0.4) Ribbon Thickness: (for helix and sheet ribbons, nucleotide ribbons, default 0.2) Protein Ribbon Width: (for helix and sheet ribbons, default 1.3) Nucleotide Ribbon Width: (for nucleotide ribbons, default 0.8) Ball Scale: (for styles 'Ball and Stick' and 'Dot', default 0.3) 4. Show Glycan Cartoon: (0: hide, 1: show, default 0) 5. Show Membrane: (0: hide, 1: show, default 1) 6. Enlarge Command Window: (0: Regular, 1: Large, default 0) Name: 1. URLs Used in Browsers Please copy one of the URLs below. They show the same result. (To add a title to share link, click "Windows > Your Note" and click "File > Share Link" again.) Original URL with commands: Lifelong Short URL:(To replace this URL, send a pull request to update share.html at iCn3D GitHub) Lifelong Short URL + Window Title:(To update the window title, click "Analysis > Your Note/Window Title".) 2. Commands Used in Jupyter Noteboook Please copy the following commands into a cell in Jupyter Notebook to show the same result. More details are at https://github.com/ncbi/icn3d/tree/master/jupyternotebook. Annotations:
Zoom: mouse wheel; Move: left button; Select Multiple Nodes: Ctrl Key and drag an Area Force on Nodes: Label Size: Internal Edges: Solvent Accessible Surface Area(SASA) calculated using the EDTSurf algorithm: (0-20% out is considered "in". 50-100% out is considered "out".) Toal: Å2 Color each residue based on the percentage of solvent accessilbe surface area. The color ranges from blue, to white, to red for a percentage of 0, 35(variable), and 100, respectively. Middle Percentage(White): % Select residue based on the percentage of solvent accessilbe surface area. The values are in the range of 0-100. Min Percentage: % Max Percentage: % Select residue based on B-factor/pLDDT. The values are in the range of 0-100. Min B-factor/pLDDT: % Max B-factor/pLDDT: % X: Y: Z: Vector 1, X: Y: Z: Vector 2, X: Y: Z: The angle is: degree. 0: 4: 8: 12: 1: 5: 9: 13: 2: 6: 10: 14: 3: 7: 11: 15: Choose an Ig template for selected residues: Choose an Ig template to align with selected residues: |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Click to View More Binding Site Information of This Target and Ligand Pair | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Click to View More Binding Site Information of This Target with Different Ligands | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Different Human System Profiles of Target | Top |
|---|---|
|
Human Similarity Proteins
of target is determined by comparing the sequence similarity of all human proteins with the target based on BLAST. The similarity proteins for a target are defined as the proteins with E-value < 0.005 and outside the protein families of the target.
A target that has fewer human similarity proteins outside its family is commonly regarded to possess a greater capacity to avoid undesired interactions and thus increase the possibility of finding successful drugs
(Brief Bioinform, 21: 649-662, 2020).
Human Tissue Distribution
of target is determined from a proteomics study that quantified more than 12,000 genes across 32 normal human tissues. Tissue Specificity (TS) score was used to define the enrichment of target across tissues.
The distribution of targets among different tissues or organs need to be taken into consideration when assessing the target druggability, as it is generally accepted that the wider the target distribution, the greater the concern over potential adverse effects
(Nat Rev Drug Discov, 20: 64-81, 2021).
Human Pathway Affiliation
of target is determined by the life-essential pathways provided on KEGG database. The target-affiliated pathways were defined based on the following two criteria (a) the pathways of the studied target should be life-essential for both healthy individuals and patients, and (b) the studied target should occupy an upstream position in the pathways and therefore had the ability to regulate biological function.
Targets involved in a fewer pathways have greater likelihood to be successfully developed, while those associated with more human pathways increase the chance of undesirable interferences with other human processes
(Pharmacol Rev, 58: 259-279, 2006).
Biological Network Descriptors
of target is determined based on a human protein-protein interactions (PPI) network consisting of 9,309 proteins and 52,713 PPIs, which were with a high confidence score of ≥ 0.95 collected from STRING database.
The network properties of targets based on protein-protein interactions (PPIs) have been widely adopted for the assessment of target’s druggability. Proteins with high node degree tend to have a high impact on network function through multiple interactions, while proteins with high betweenness centrality are regarded to be central for communication in interaction networks and regulate the flow of signaling information
(Front Pharmacol, 9, 1245, 2018;
Curr Opin Struct Biol. 44:134-142, 2017).
Human Similarity Proteins
Human Tissue Distribution
Human Pathway Affiliation
Biological Network Descriptors
|
|
|
Note:
If a protein has TS (tissue specficity) scores at least in one tissue >= 2.5, this protein is called tissue-enriched (including tissue-enriched-but-not-specific and tissue-specific). In the plots, the vertical lines are at thresholds 2.5 and 4.
|
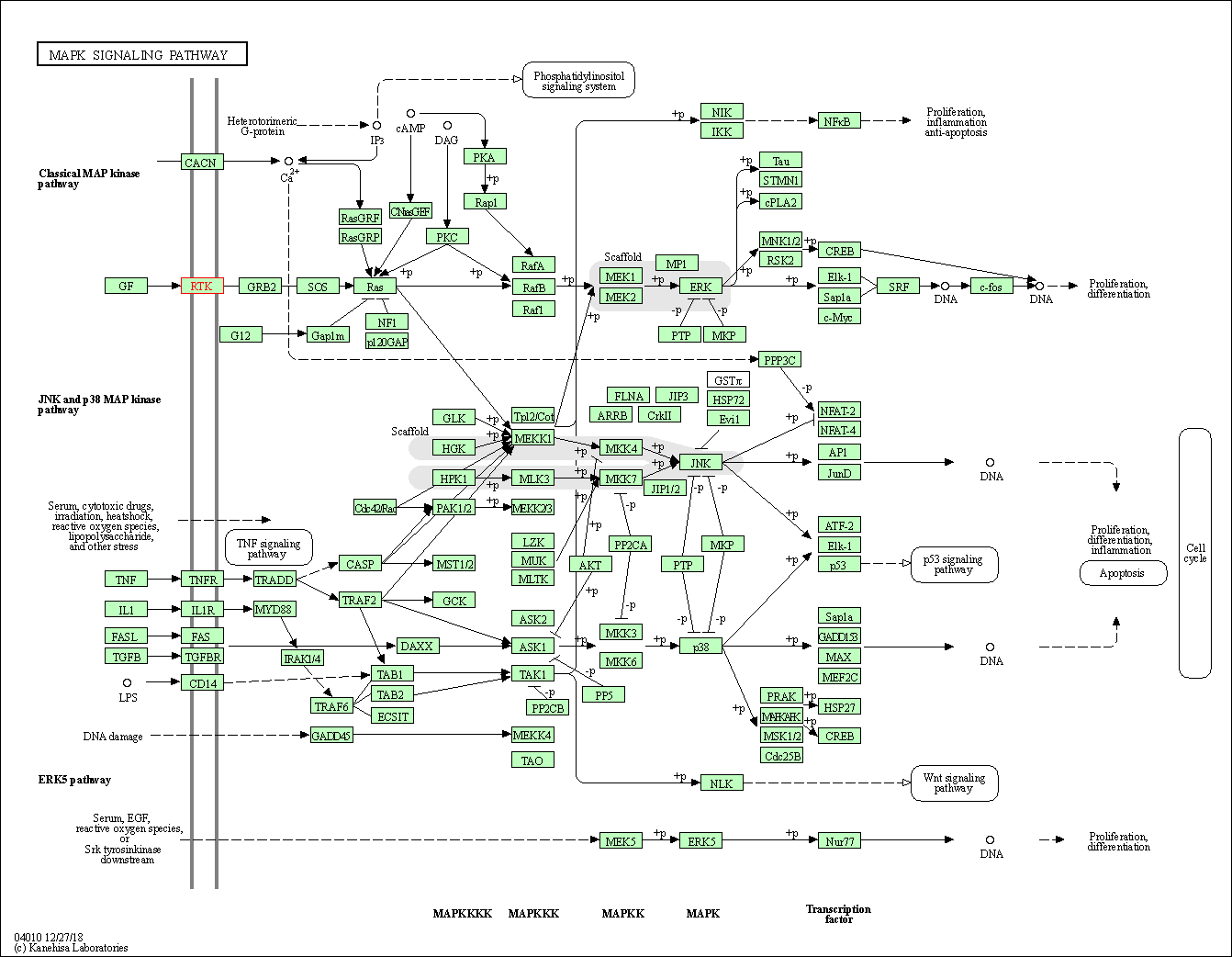
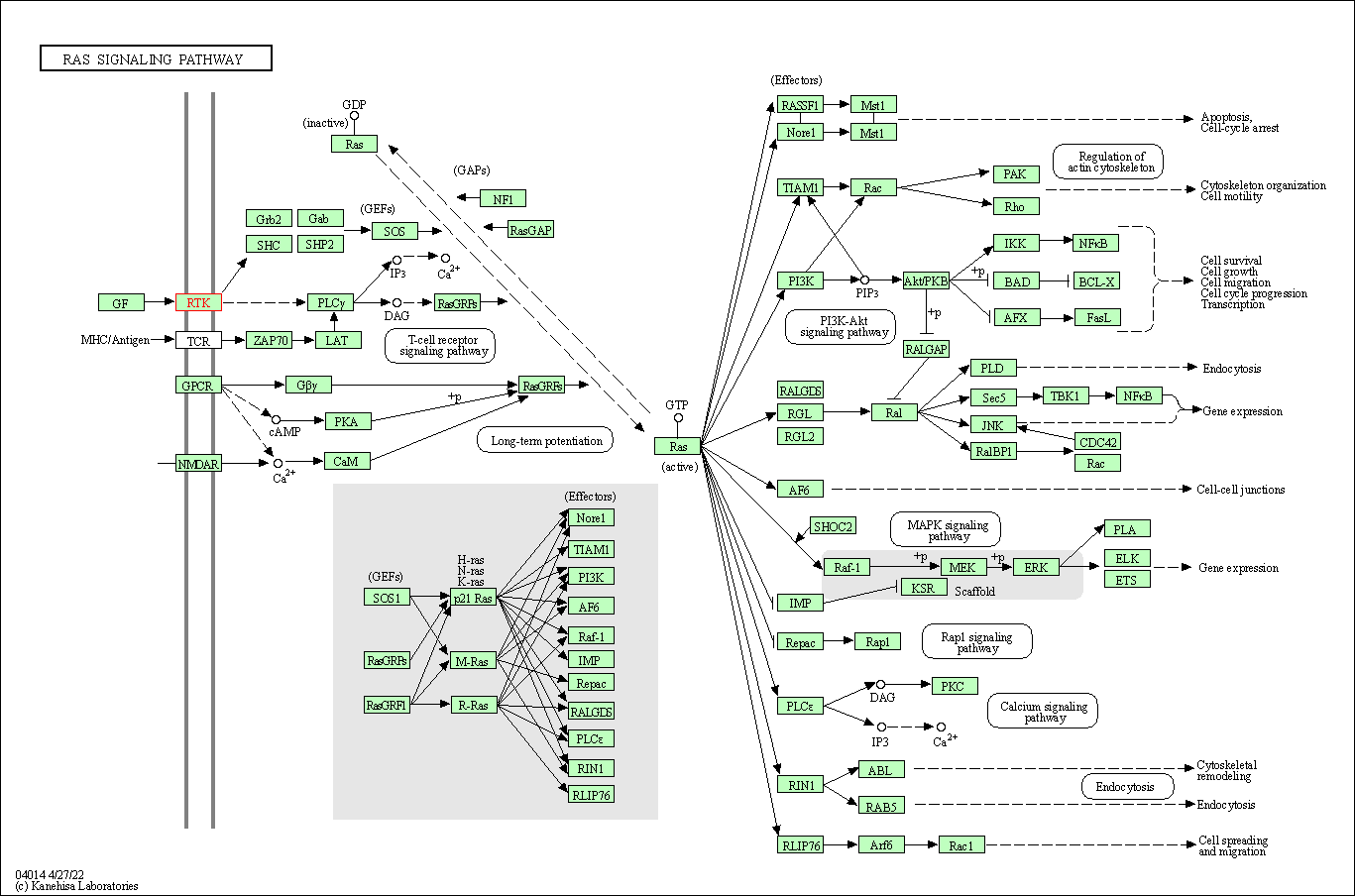
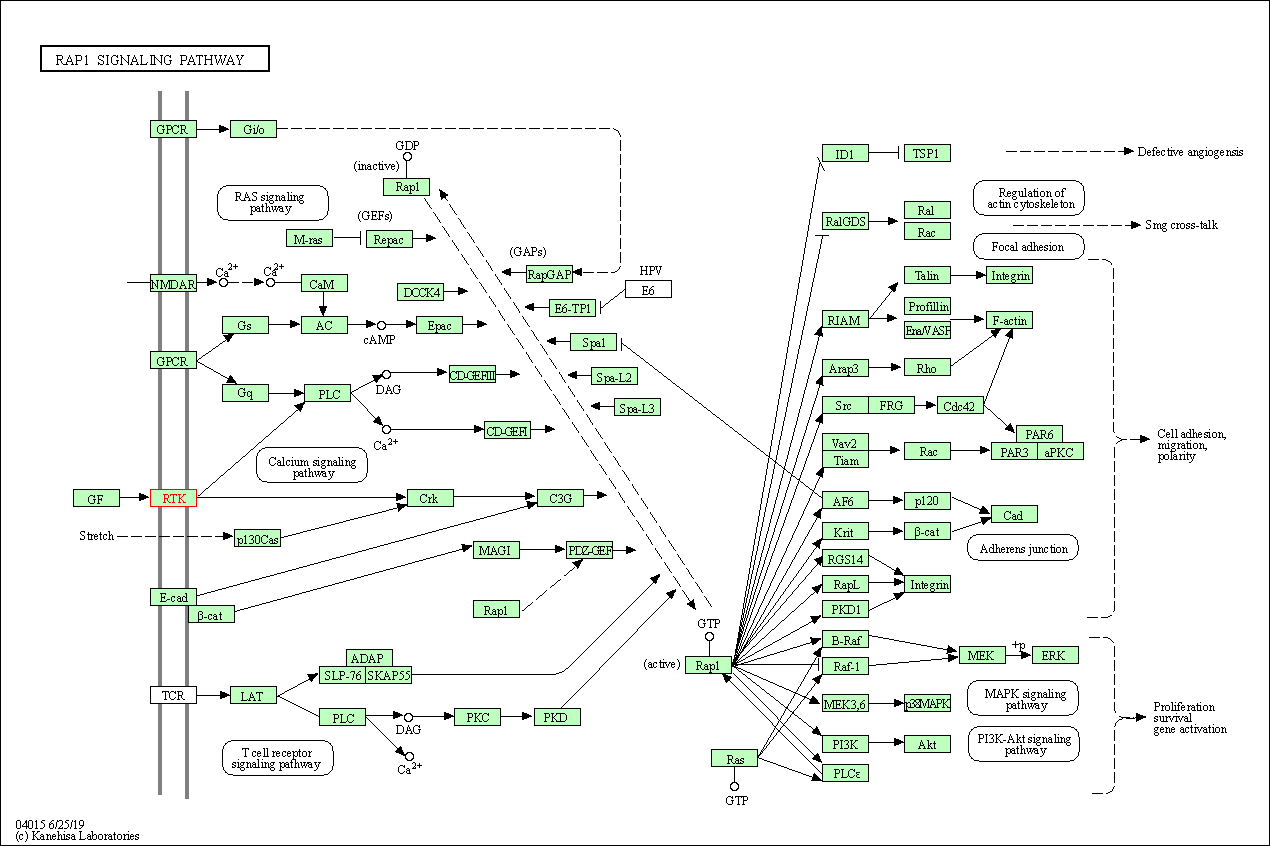
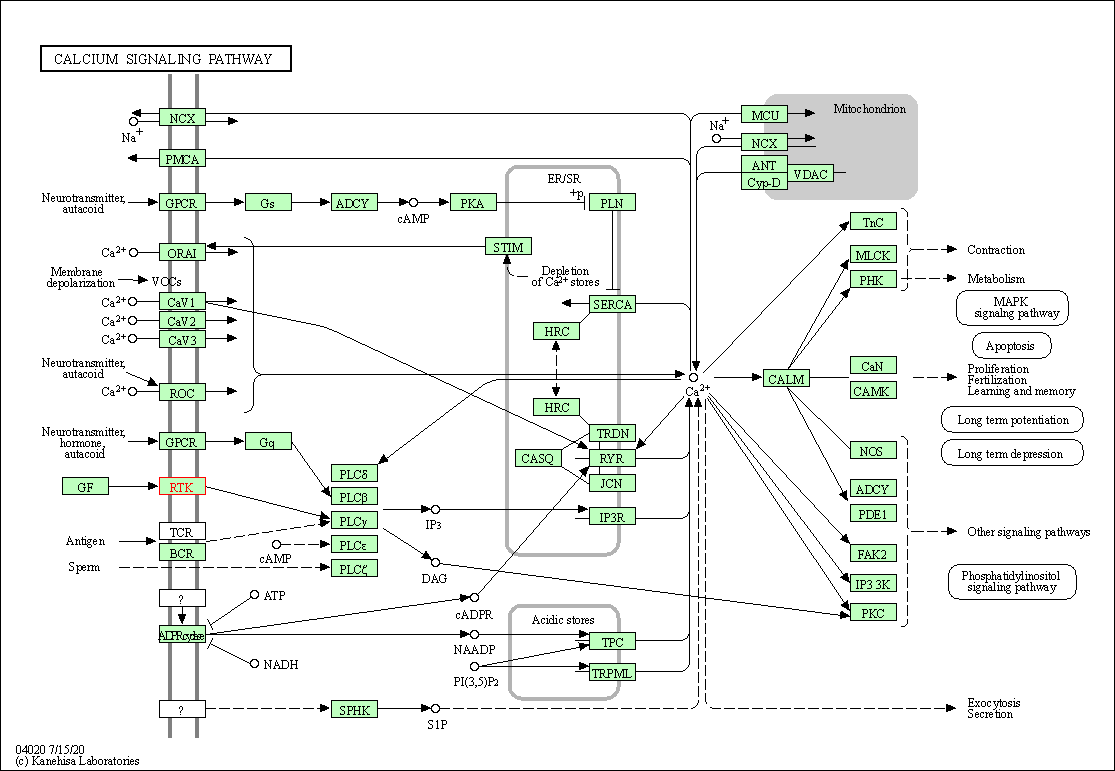
| KEGG Pathway | Pathway ID | Affiliated Target | Pathway Map |
|---|---|---|---|
| MAPK signaling pathway | hsa04010 | Affiliated Target |

|
| Class: Environmental Information Processing => Signal transduction | Pathway Hierarchy | ||
| Ras signaling pathway | hsa04014 | Affiliated Target |

|
| Class: Environmental Information Processing => Signal transduction | Pathway Hierarchy | ||
| Rap1 signaling pathway | hsa04015 | Affiliated Target |

|
| Class: Environmental Information Processing => Signal transduction | Pathway Hierarchy | ||
| Calcium signaling pathway | hsa04020 | Affiliated Target |

|
| Class: Environmental Information Processing => Signal transduction | Pathway Hierarchy | ||
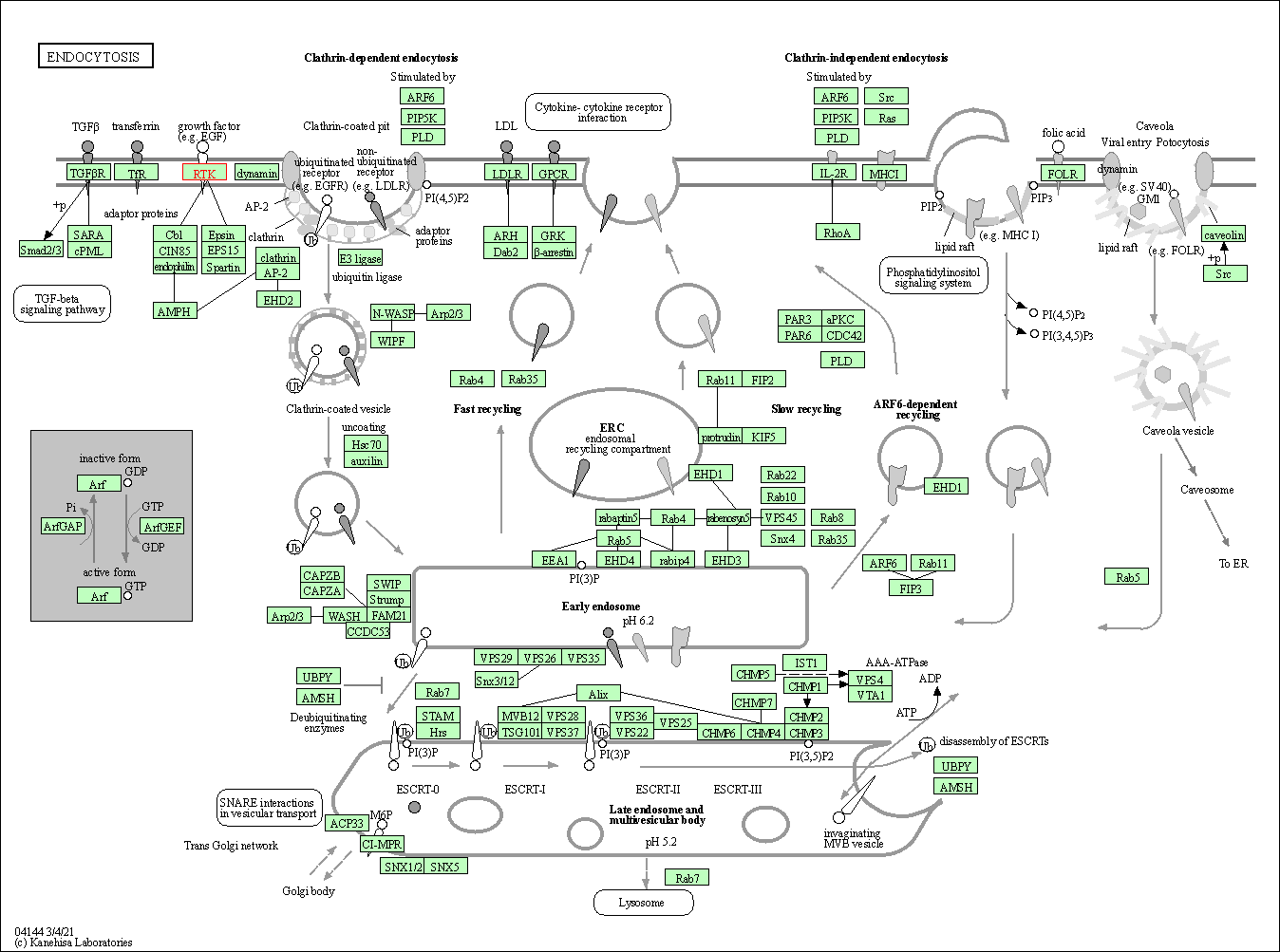
| Endocytosis | hsa04144 | Affiliated Target |

|
| Class: Cellular Processes => Transport and catabolism | Pathway Hierarchy | ||
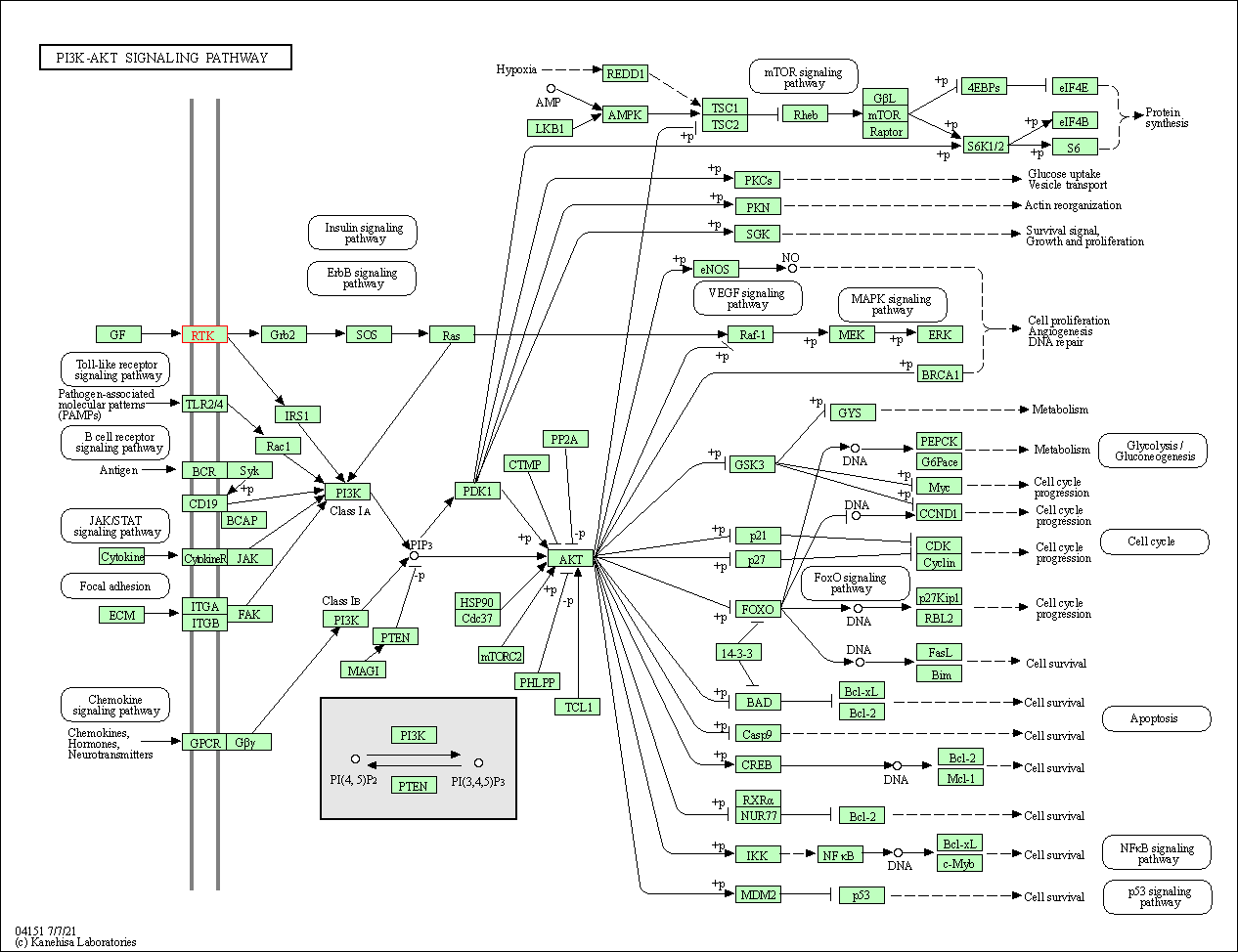
| PI3K-Akt signaling pathway | hsa04151 | Affiliated Target |

|
| Class: Environmental Information Processing => Signal transduction | Pathway Hierarchy | ||
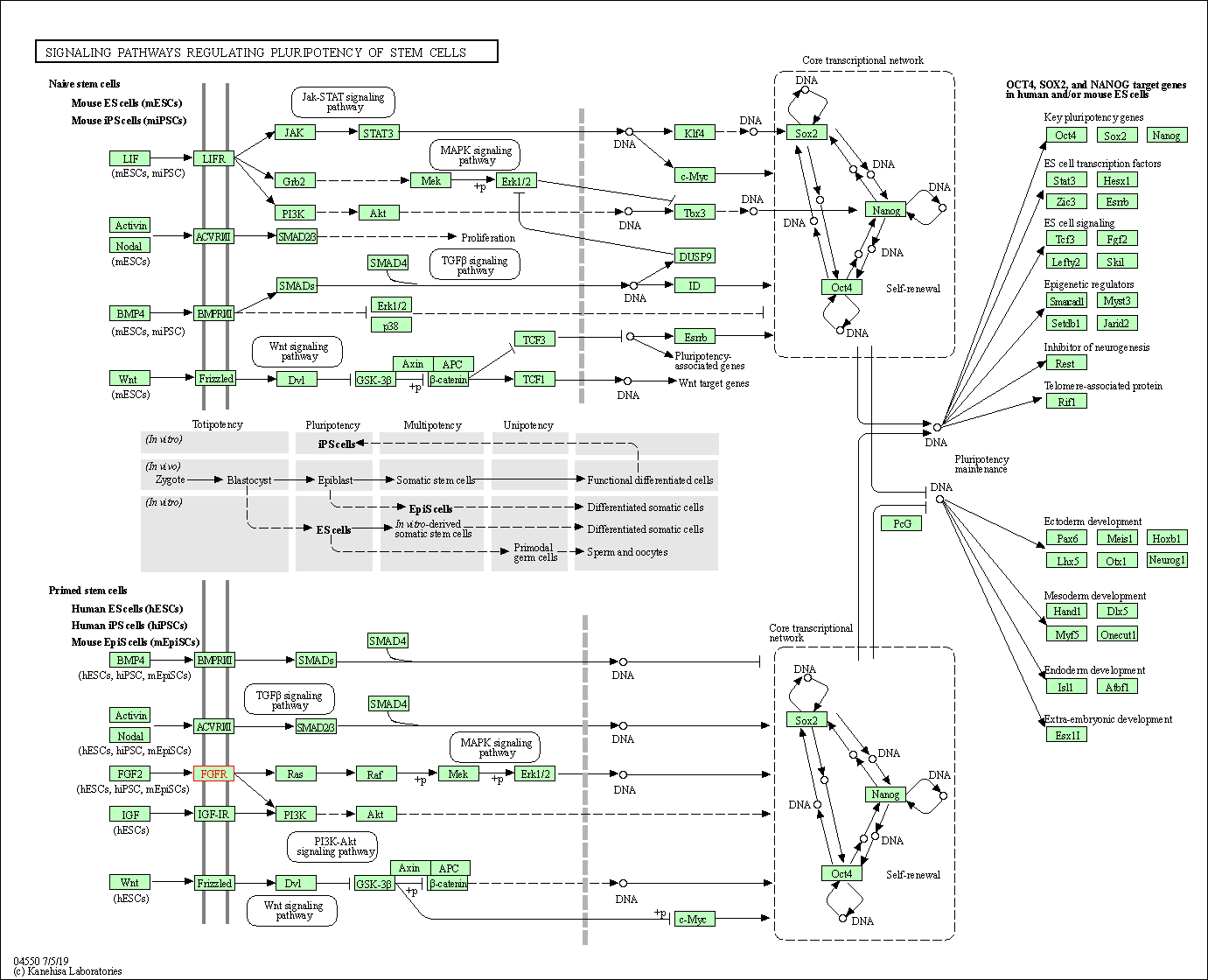
| Signaling pathways regulating pluripotency of stem cells | hsa04550 | Affiliated Target |

|
| Class: Cellular Processes => Cellular community - eukaryotes | Pathway Hierarchy | ||
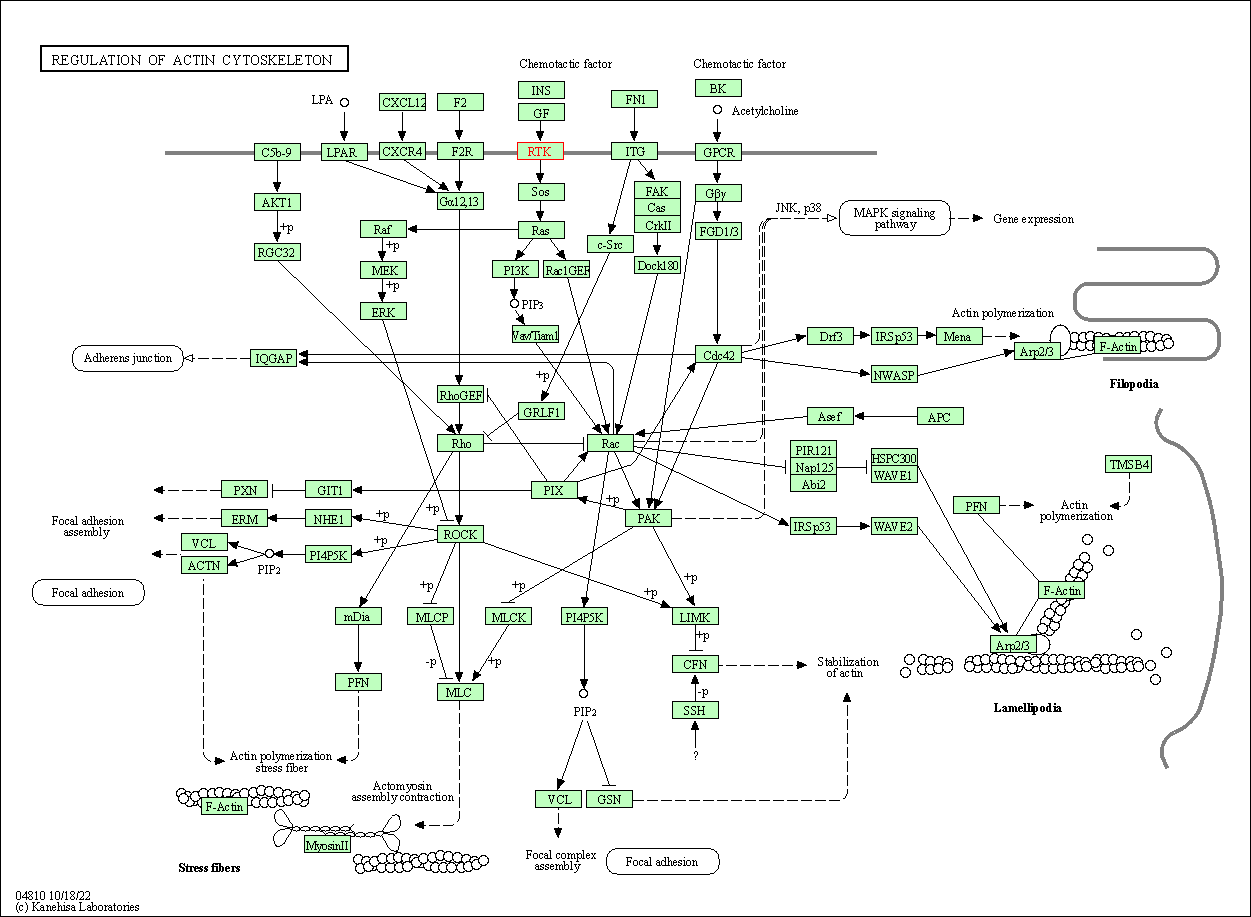
| Regulation of actin cytoskeleton | hsa04810 | Affiliated Target |

|
| Class: Cellular Processes => Cell motility | Pathway Hierarchy | ||
| Click to Show/Hide the Information of Affiliated Human Pathways | |||
| Degree | 20 | Degree centrality | 2.15E-03 | Betweenness centrality | 6.13E-04 |
|---|---|---|---|---|---|
| Closeness centrality | 2.35E-01 | Radiality | 1.41E+01 | Clustering coefficient | 2.74E-01 |
| Neighborhood connectivity | 4.63E+01 | Topological coefficient | 1.12E-01 | Eccentricity | 12 |
| Download | Click to Download the Full PPI Network of This Target | ||||
| Chemical Structure based Activity Landscape of Target | Top |
|---|---|
| Drug Property Profile of Target | Top | |
|---|---|---|
| (1) Molecular Weight (mw) based Drug Clustering | (2) Octanol/Water Partition Coefficient (xlogp) based Drug Clustering | |
|
|
||
| (3) Hydrogen Bond Donor Count (hbonddonor) based Drug Clustering | (4) Hydrogen Bond Acceptor Count (hbondacc) based Drug Clustering | |
|
|
||
| (5) Rotatable Bond Count (rotbonds) based Drug Clustering | (6) Topological Polar Surface Area (polararea) based Drug Clustering | |
|
|
||
| "RO5" indicates the cutoff set by lipinski's rule of five; "D123AB" colored in GREEN denotes the no violation of any cutoff in lipinski's rule of five; "D123AB" colored in PURPLE refers to the violation of only one cutoff in lipinski's rule of five; "D123AB" colored in BLACK represents the violation of more than one cutoffs in lipinski's rule of five | ||
| Co-Targets | Top | |||||
|---|---|---|---|---|---|---|
| Co-Targets | ||||||
| Target Poor or Non Binders | Top | |||||
|---|---|---|---|---|---|---|
| Target Poor or Non Binders | ||||||
| Target Regulators | Top | |||||
|---|---|---|---|---|---|---|
| Target-regulating microRNAs | ||||||
| Target-interacting Proteins | ||||||
| Target Profiles in Patients | Top | |||||
|---|---|---|---|---|---|---|
| Target Expression Profile (TEP) |
||||||
| Drug Resistance Mutation (DRM) |
||||||
| Target Affiliated Biological Pathways | Top | |||||
|---|---|---|---|---|---|---|
| KEGG Pathway | [+] 11 KEGG Pathways | + | ||||
| 1 | MAPK signaling pathway | |||||
| 2 | Ras signaling pathway | |||||
| 3 | Rap1 signaling pathway | |||||
| 4 | Endocytosis | |||||
| 5 | PI3K-Akt signaling pathway | |||||
| 6 | Signaling pathways regulating pluripotency of stem cells | |||||
| 7 | Regulation of actin cytoskeleton | |||||
| 8 | Pathways in cancer | |||||
| 9 | MicroRNAs in cancer | |||||
| 10 | Bladder cancer | |||||
| 11 | Central carbon metabolism in cancer | |||||
| Panther Pathway | [+] 1 Panther Pathways | + | ||||
| 1 | FGF signaling pathway | |||||
| Reactome | [+] 1 Reactome Pathways | + | ||||
| 1 | FGFR3 mutant receptor activation | |||||
| WikiPathways | [+] 5 WikiPathways | + | ||||
| 1 | Regulation of Actin Cytoskeleton | |||||
| 2 | Endochondral Ossification | |||||
| 3 | Bladder Cancer | |||||
| 4 | Neural Crest Differentiation | |||||
| 5 | Signaling by FGFR | |||||
| Target-Related Models and Studies | Top | |||||
|---|---|---|---|---|---|---|
| Target Validation | ||||||
| References | Top | |||||
|---|---|---|---|---|---|---|
| REF 1 | Emerging therapies for multiple myeloma. Expert Opin Emerg Drugs. 2009 Mar;14(1):99-127. | |||||
| REF 2 | Drugs@FDA. U.S. Food and Drug Administration. U.S. Department of Health Human Services. 2020 | |||||
| REF 3 | ClinicalTrials.gov (NCT00914420) Optical Coherence Tomography (OCT) Evaluation of Re-endothelization: A Comparison of the Intrepide Stent Versus Taxus . U.S. National Institutes of Health. | |||||
| REF 4 | URL: http://www.guidetopharmacology.org Nucleic Acids Res. 2015 Oct 12. pii: gkv1037. The IUPHAR/BPS Guide to PHARMACOLOGY in 2016: towards curated quantitative interactions between 1300 protein targets and 6000 ligands. (Ligand id: 8097). | |||||
| REF 5 | ClinicalTrials.gov (NCT00908752) Phase III Trans-Arterial Chemo-Embolization (TACE) Adjuvant HCC. U.S. National Institutes of Health. | |||||
| REF 6 | URL: http://www.guidetopharmacology.org Nucleic Acids Res. 2015 Oct 12. pii: gkv1037. The IUPHAR/BPS Guide to PHARMACOLOGY in 2016: towards curated quantitative interactions between 1300 protein targets and 6000 ligands. (Ligand id: 7649). | |||||
| REF 7 | ClinicalTrials.gov (NCT02135107) A Double-blind Comparative Study of the Efficacy and Safety of E3810 10mg Once and Twice Daily in Maintenance Therapy for PPI Resistant Gastroesophageal Reflux Disease Patients. U.S. National Institutes of Health. | |||||
| REF 8 | ClinicalTrials.gov (NCT01223027) Study of Dovitinib Versus Sorafenib in Patients With Metastatic Renal Cell Carcinoma. U.S. National Institutes of Health. | |||||
| REF 9 | Clinical pipeline report, company report or official report of the Pharmaceutical Research and Manufacturers of America (PhRMA) | |||||
| REF 10 | Clinical pipeline report, company report or official report of the Pharmaceutical Research and Manufacturers of America (PhRMA) | |||||
| REF 11 | ClinicalTrials.gov (NCT03834220) Basket Trial in Solid Tumors Harboring a Fusion of FGFR1, FGFR2 or FGFR3- (FUZE Clinical Trial). U.S. National Institutes of Health. | |||||
| REF 12 | ClinicalTrials.gov (NCT04638153) A Study Of Safety, Tolerability And Effectiveness Of Recifercept In Children With Achondroplasia. U.S. National Institutes of Health. | |||||
| REF 13 | URL: http://www.guidetopharmacology.org Nucleic Acids Res. 2015 Oct 12. pii: gkv1037. The IUPHAR/BPS Guide to PHARMACOLOGY in 2016: towards curated quantitative interactions between 1300 protein targets and 6000 ligands. (Ligand id: 7643). | |||||
| REF 14 | ClinicalTrials.gov (NCT00107237) AEE788 and Everolimus in Treating Patients With Recurrent or Relapsed Glioblastoma Multiforme. U.S. National Institutes of Health. | |||||
| REF 15 | MK-2461, a novel multitargeted kinase inhibitor, preferentially inhibits the activated c-Met receptor. Cancer Res. 2010 Feb 15;70(4):1524-33. | |||||
| REF 16 | Clinical pipeline report, company report or official report of Genentech (2011). | |||||
| REF 17 | Clinical pipeline report, company report or official report of Sanofi | |||||
| REF 18 | Trusted, scientifically sound profiles of drug programs, clinical trials, safety reports, and company deals, written by scientists. Springer. 2015. Adis Insight (drug id 800014130) | |||||
| REF 19 | A comparison of physicochemical property profiles of marketed oral drugs and orally bioavailable anti-cancer protein kinase inhibitors in clinical development. Curr Top Med Chem. 2007;7(14):1408-22. | |||||
| REF 20 | E-3810 is a potent dual inhibitor of VEGFR and FGFR that exerts antitumor activity in multiple preclinical models. Cancer Res. 2011 Feb 15;71(4):1396-405. | |||||
| REF 21 | In vitro and in vivo characterization of Recifercept, a soluble fibroblast growth factor receptor 3, as treatment for achondroplasia. PLoS One. 2020 Dec 28;15(12):e0244368. | |||||
| REF 22 | Pyrido[2,3-d]pyrimidin-7-one inhibitors of cyclin-dependent kinases. J Med Chem. 2000 Nov 30;43(24):4606-16. | |||||
| REF 23 | Molecular modeling of wild-type and D816V c-Kit inhibition based on ATP-competitive binding of ellipticine derivatives to tyrosine kinases. J Med Chem. 2005 Oct 6;48(20):6194-201. | |||||
| REF 24 | URL: http://www.guidetopharmacology.org Nucleic Acids Res. 2015 Oct 12. pii: gkv1037. The IUPHAR/BPS Guide to PHARMACOLOGY in 2016: towards curated quantitative interactions between 1300 protein targets and 6000 ligands. (Target id: 1810). | |||||
| REF 25 | In vitro and in vivo evaluation of 6-aminopyrazolyl-pyridine-3-carbonitriles as JAK2 kinase inhibitors. Bioorg Med Chem Lett. 2011 May 15;21(10):2958-61. | |||||
| REF 26 | Biological evaluation of a multi-targeted small molecule inhibitor of tumor-induced angiogenesis. Bioorg Med Chem Lett. 2006 Apr 1;16(7):1950-3. | |||||
| REF 27 | Molecular basis for receptor tyrosine kinase A-loop tyrosine transphosphorylation. Nat Chem Biol. 2020 Mar;16(3):267-277. | |||||
| REF 28 | Structure-based drug design of 1,3,5-triazine and pyrimidine derivatives as novel FGFR3 inhibitors with high selectivity over VEGFR2. Bioorg Med Chem. 2020 May 15;28(10):115453. | |||||
If You Find Any Error in Data or Bug in Web Service, Please Kindly Report It to Dr. Zhou and Dr. Zhang.

