Target Information
| Target General Information | Top | |||||
|---|---|---|---|---|---|---|
| Target ID |
T59190
(Former ID: TTDR00557)
|
|||||
| Target Name |
Protein kinase C zeta (PRKCZ)
|
|||||
| Synonyms |
Protein kinase C zeta type; PKC2; NPKC-zeta
Click to Show/Hide
|
|||||
| Gene Name |
PRKCZ
|
|||||
| Target Type |
Patented-recorded target
|
[1] | ||||
| Disease | [+] 1 Target-related Diseases | + | ||||
| 1 | Solid tumour/cancer [ICD-11: 2A00-2F9Z] | |||||
| Function |
Upon lipopolysaccharide (LPS) treatment in macrophages, or following mitogenic stimuli, functions downstream of PI3K to activate MAP2K1/MEK1-MAPK1/ERK2 signaling cascade independently of RAF1 activation. Required for insulin-dependent activation of AKT3, but may function as an adapter rather than a direct activator. Upon insulin treatment may act as a downstream effector of PI3K and contribute to the activation of translocation of the glucose transporter SLC2A4/GLUT4 and subsequent glucose transport in adipocytes. In EGF-induced cells, binds and activates MAP2K5/MEK5-MAPK7/ERK5 independently of its kinase activity and can activate JUN promoter through MEF2C. Through binding with SQSTM1/p62, functions in interleukin-1 signaling and activation of NF-kappa-B with the specific adapters RIPK1 and TRAF6. Participates in TNF-dependent transactivation of NF-kappa-B by phosphorylating and activating IKBKB kinase, which in turn leads to the degradation of NF-kappa-B inhibitors. In migrating astrocytes, forms a cytoplasmic complex with PARD6A and is recruited by CDC42 to function in the establishment of cell polarity along with the microtubule motor and dynein. In association with FEZ1, stimulates neuronal differentiation in PC12 cells. In the inflammatory response, is required for the T-helper 2 (Th2) differentiation process, including interleukin production, efficient activation of JAK1 and the subsequent phosphorylation and nuclear translocation of STAT6. May be involved in development of allergic airway inflammation (asthma), a process dependent on Th2 immune response. In the NF-kappa-B-mediated inflammatory response, can relieve SETD6-dependent repression of NF-kappa-B target genes by phosphorylating the RELA subunit at 'Ser-311'. In vein endothelial cells treated with the oxidant peroxynitrite, phosphorylates STK11 leading to nuclear export of STK11, subsequent inhibition of PI3K/Akt signaling, and increased apoptosis. Phosphorylates VAMP2 in vitro. Calcium- and diacylglycerol-independent serine/threonine-protein kinase that functions in phosphatidylinositol 3-kinase (PI3K) pathway and mitogen-activated protein (MAP) kinase cascade, and is involved in NF-kappa-B activation, mitogenic signaling, cell proliferation, cell polarity, inflammatory response and maintenance of long-term potentiation (LTP).
Click to Show/Hide
|
|||||
| BioChemical Class |
Kinase
|
|||||
| UniProt ID | ||||||
| EC Number |
EC 2.7.11.13
|
|||||
| Sequence |
MPSRTGPKMEGSGGRVRLKAHYGGDIFITSVDAATTFEELCEEVRDMCRLHQQHPLTLKW
VDSEGDPCTVSSQMELEEAFRLARQCRDEGLIIHVFPSTPEQPGLPCPGEDKSIYRRGAR RWRKLYRANGHLFQAKRFNRRAYCGQCSERIWGLARQGYRCINCKLLVHKRCHGLVPLTC RKHMDSVMPSQEPPVDDKNEDADLPSEETDGIAYISSSRKHDSIKDDSEDLKPVIDGMDG IKISQGLGLQDFDLIRVIGRGSYAKVLLVRLKKNDQIYAMKVVKKELVHDDEDIDWVQTE KHVFEQASSNPFLVGLHSCFQTTSRLFLVIEYVNGGDLMFHMQRQRKLPEEHARFYAAEI CIALNFLHERGIIYRDLKLDNVLLDADGHIKLTDYGMCKEGLGPGDTTSTFCGTPNYIAP EILRGEEYGFSVDWWALGVLMFEMMAGRSPFDIITDNPDMNTEDYLFQVILEKPIRIPRF LSVKASHVLKGFLNKDPKERLGCRPQTGFSDIKSHAFFRSIDWDLLEKKQALPPFQPQIT DDYGLDNFDTQFTSEPVQLTPDDEDAIKRIDQSEFEGFEYINPLLLSTEESV Click to Show/Hide
|
|||||
| 3D Structure | Click to Show 3D Structure of This Target | AlphaFold | ||||
| HIT2.0 ID | T34FQX | |||||
| Cell-based Target Expression Variations | Top | |||||
|---|---|---|---|---|---|---|
| Cell-based Target Expression Variations | ||||||
| Different Human System Profiles of Target | Top |
|---|---|
|
Human Similarity Proteins
of target is determined by comparing the sequence similarity of all human proteins with the target based on BLAST. The similarity proteins for a target are defined as the proteins with E-value < 0.005 and outside the protein families of the target.
A target that has fewer human similarity proteins outside its family is commonly regarded to possess a greater capacity to avoid undesired interactions and thus increase the possibility of finding successful drugs
(Brief Bioinform, 21: 649-662, 2020).
Human Tissue Distribution
of target is determined from a proteomics study that quantified more than 12,000 genes across 32 normal human tissues. Tissue Specificity (TS) score was used to define the enrichment of target across tissues.
The distribution of targets among different tissues or organs need to be taken into consideration when assessing the target druggability, as it is generally accepted that the wider the target distribution, the greater the concern over potential adverse effects
(Nat Rev Drug Discov, 20: 64-81, 2021).
Human Pathway Affiliation
of target is determined by the life-essential pathways provided on KEGG database. The target-affiliated pathways were defined based on the following two criteria (a) the pathways of the studied target should be life-essential for both healthy individuals and patients, and (b) the studied target should occupy an upstream position in the pathways and therefore had the ability to regulate biological function.
Targets involved in a fewer pathways have greater likelihood to be successfully developed, while those associated with more human pathways increase the chance of undesirable interferences with other human processes
(Pharmacol Rev, 58: 259-279, 2006).
Biological Network Descriptors
of target is determined based on a human protein-protein interactions (PPI) network consisting of 9,309 proteins and 52,713 PPIs, which were with a high confidence score of ≥ 0.95 collected from STRING database.
The network properties of targets based on protein-protein interactions (PPIs) have been widely adopted for the assessment of target’s druggability. Proteins with high node degree tend to have a high impact on network function through multiple interactions, while proteins with high betweenness centrality are regarded to be central for communication in interaction networks and regulate the flow of signaling information
(Front Pharmacol, 9, 1245, 2018;
Curr Opin Struct Biol. 44:134-142, 2017).
Human Similarity Proteins
Human Tissue Distribution
Human Pathway Affiliation
Biological Network Descriptors
|
|
|
Note:
If a protein has TS (tissue specficity) scores at least in one tissue >= 2.5, this protein is called tissue-enriched (including tissue-enriched-but-not-specific and tissue-specific). In the plots, the vertical lines are at thresholds 2.5 and 4.
|
| KEGG Pathway | Pathway ID | Affiliated Target | Pathway Map |
|---|---|---|---|
| Rap1 signaling pathway | hsa04015 | Affiliated Target |
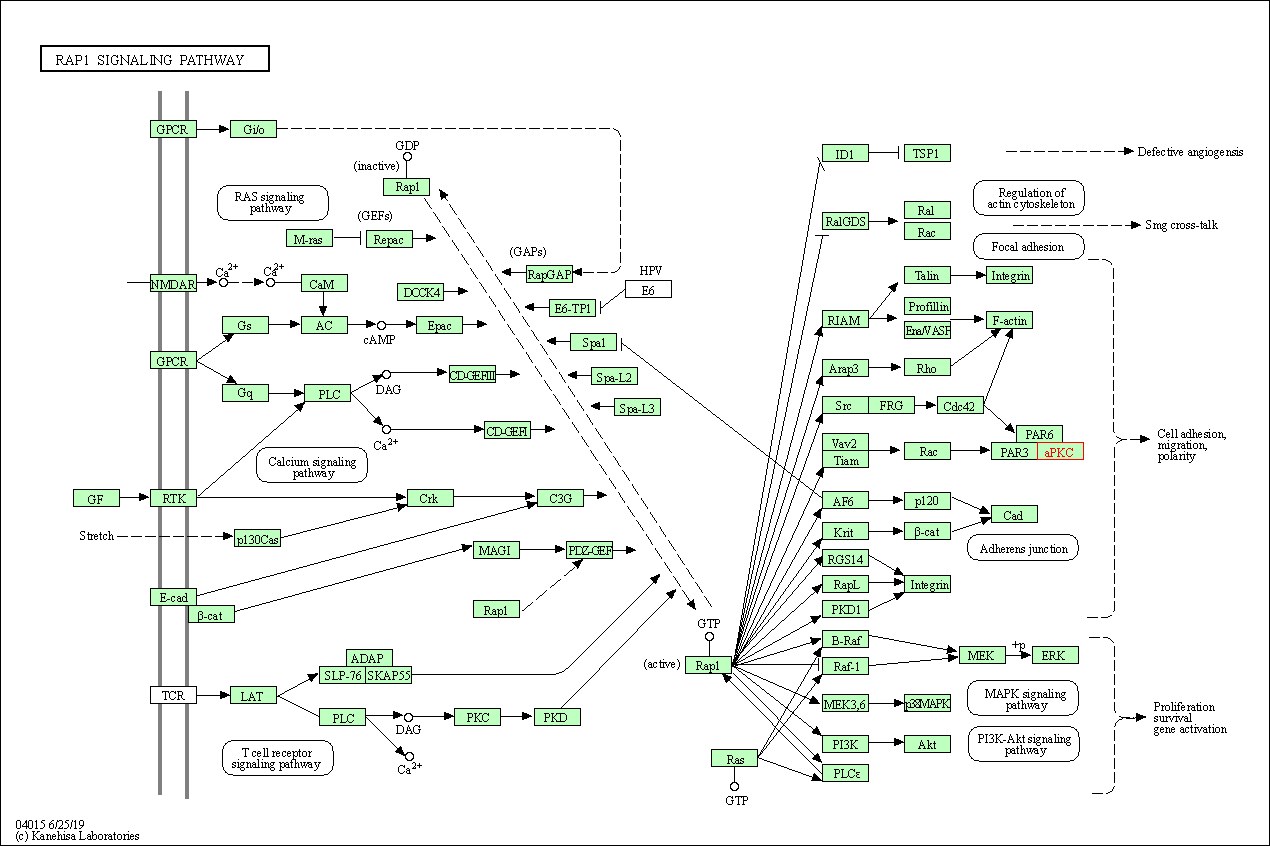
|
| Class: Environmental Information Processing => Signal transduction | Pathway Hierarchy | ||
| Chemokine signaling pathway | hsa04062 | Affiliated Target |

|
| Class: Organismal Systems => Immune system | Pathway Hierarchy | ||
| Sphingolipid signaling pathway | hsa04071 | Affiliated Target |
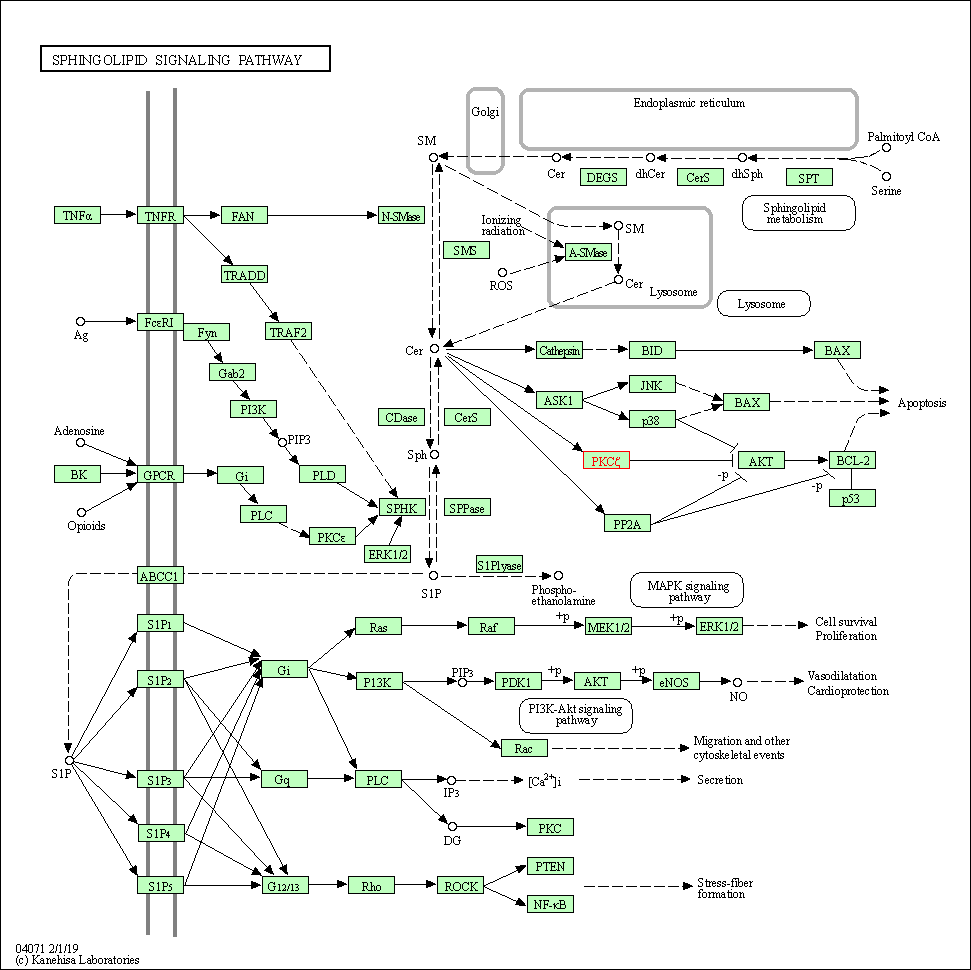
|
| Class: Environmental Information Processing => Signal transduction | Pathway Hierarchy | ||
| Endocytosis | hsa04144 | Affiliated Target |
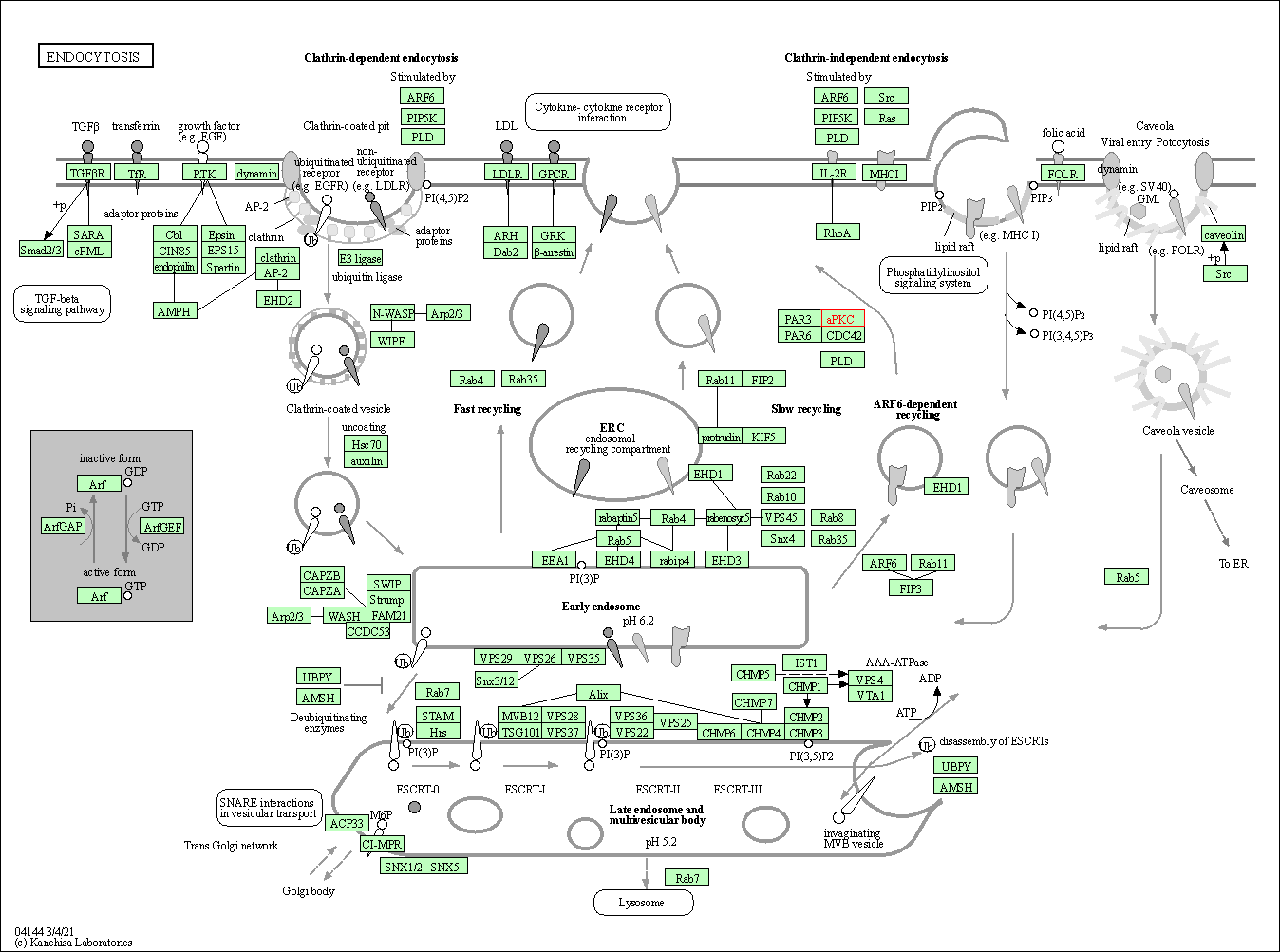
|
| Class: Cellular Processes => Transport and catabolism | Pathway Hierarchy | ||
| Axon guidance | hsa04360 | Affiliated Target |
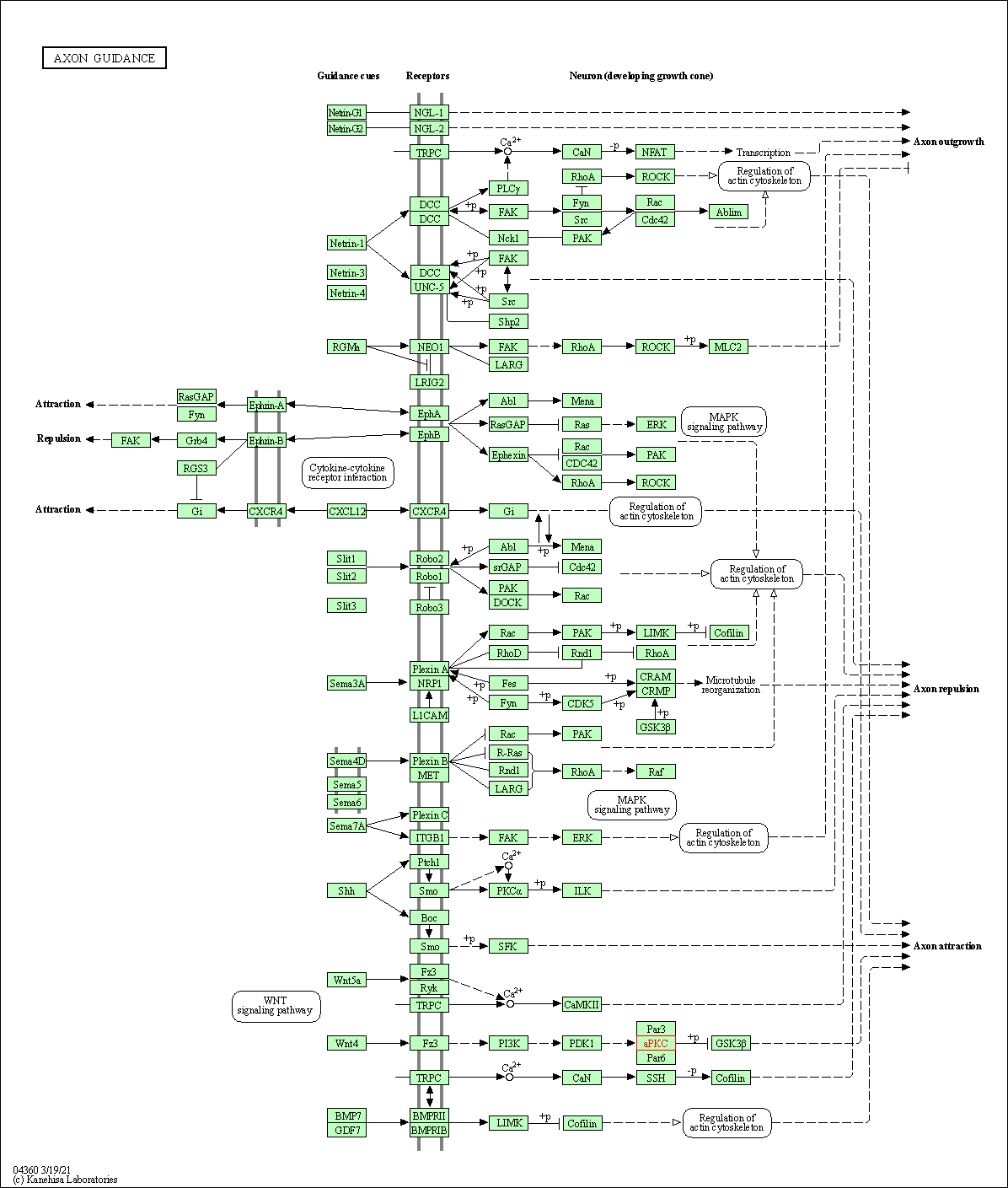
|
| Class: Organismal Systems => Development and regeneration | Pathway Hierarchy | ||
| Hippo signaling pathway | hsa04390 | Affiliated Target |
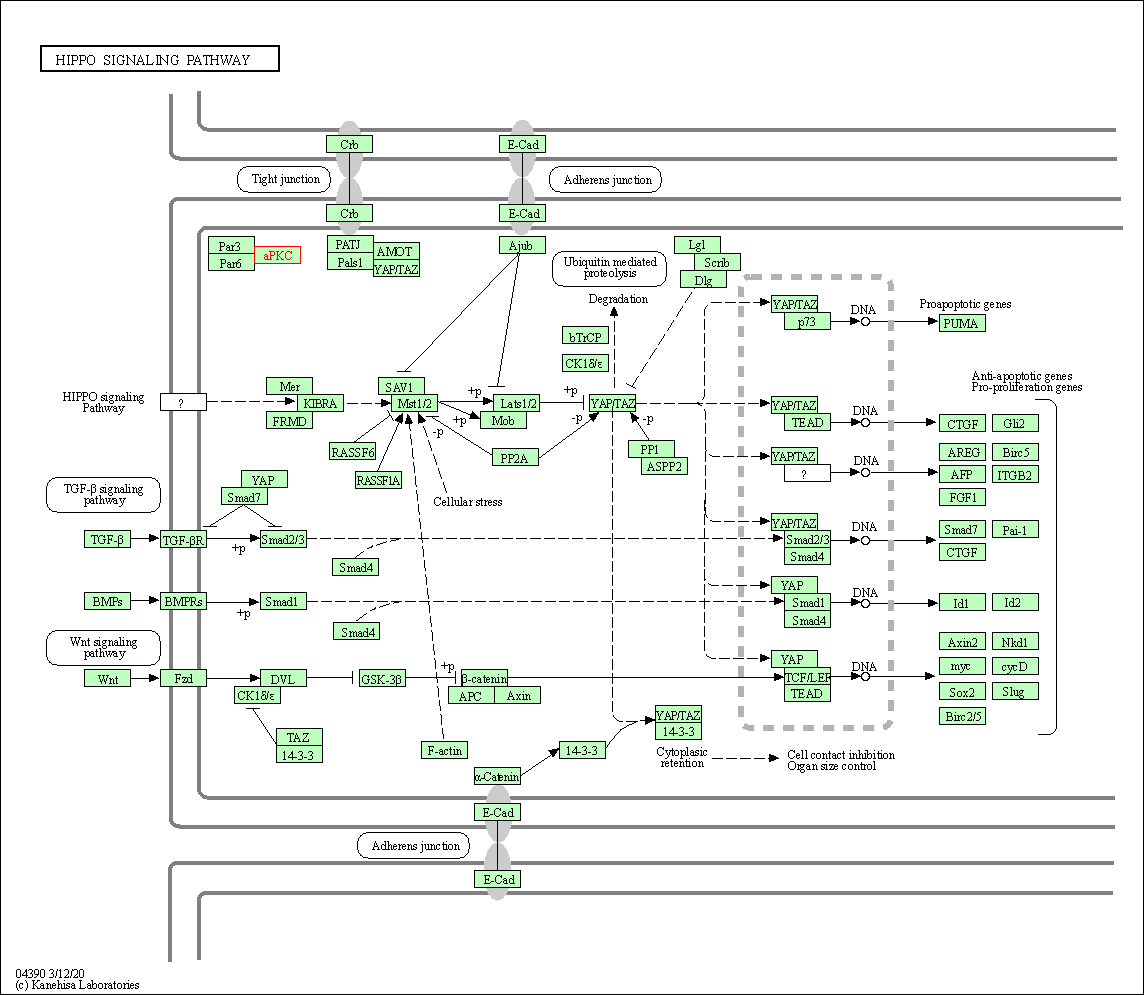
|
| Class: Environmental Information Processing => Signal transduction | Pathway Hierarchy | ||
| Tight junction | hsa04530 | Affiliated Target |
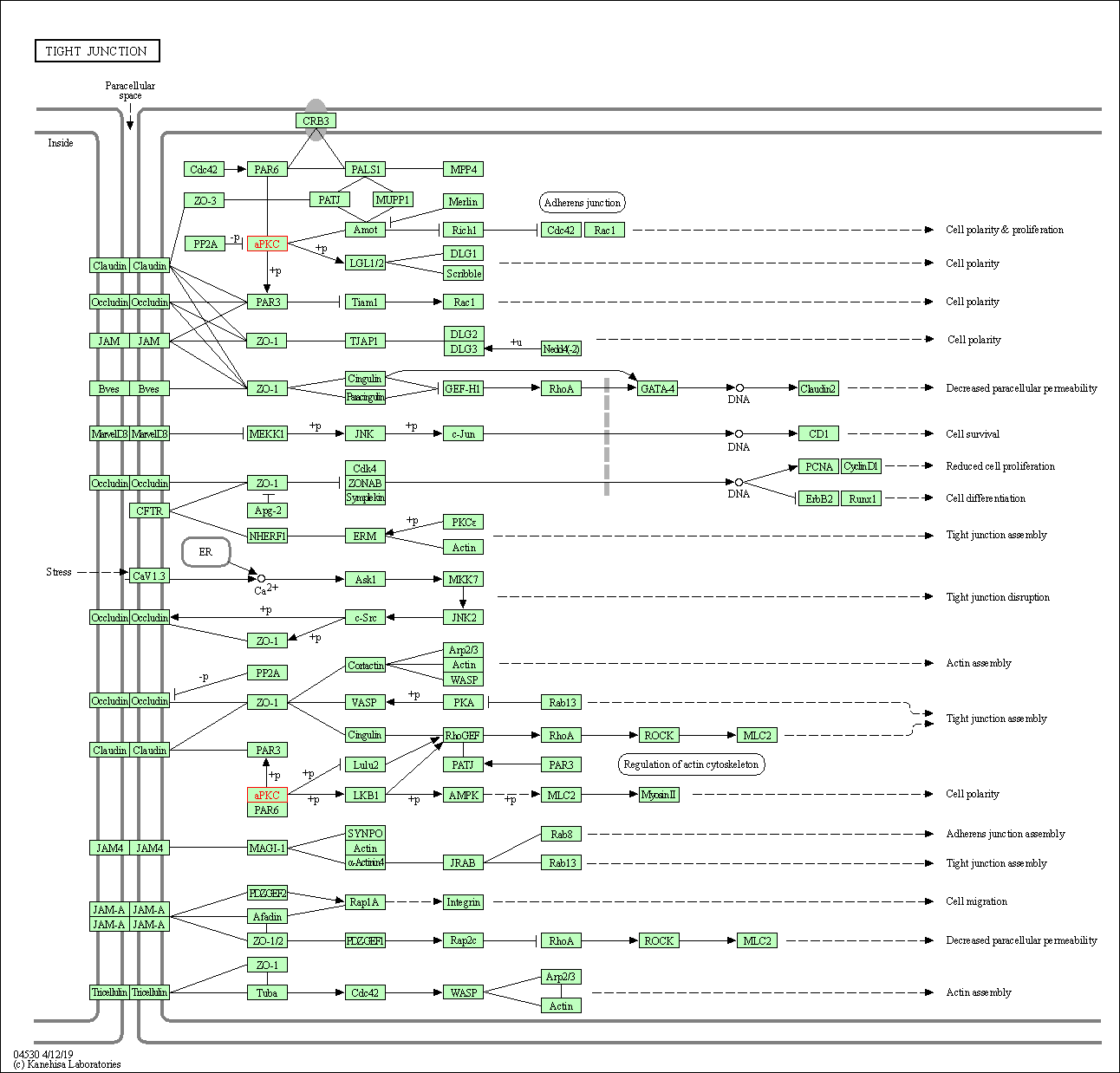
|
| Class: Cellular Processes => Cellular community - eukaryotes | Pathway Hierarchy | ||
| Platelet activation | hsa04611 | Affiliated Target |

|
| Class: Organismal Systems => Immune system | Pathway Hierarchy | ||
| Insulin signaling pathway | hsa04910 | Affiliated Target |

|
| Class: Organismal Systems => Endocrine system | Pathway Hierarchy | ||
| Relaxin signaling pathway | hsa04926 | Affiliated Target |

|
| Class: Organismal Systems => Endocrine system | Pathway Hierarchy | ||
| Click to Show/Hide the Information of Affiliated Human Pathways | |||
| Degree | 24 | Degree centrality | 2.58E-03 | Betweenness centrality | 9.67E-04 |
|---|---|---|---|---|---|
| Closeness centrality | 2.56E-01 | Radiality | 1.44E+01 | Clustering coefficient | 1.81E-01 |
| Neighborhood connectivity | 4.80E+01 | Topological coefficient | 6.50E-02 | Eccentricity | 11 |
| Download | Click to Download the Full PPI Network of This Target | ||||
| Chemical Structure based Activity Landscape of Target | Top |
|---|---|
| Drug Property Profile of Target | Top | |
|---|---|---|
| (1) Molecular Weight (mw) based Drug Clustering | (2) Octanol/Water Partition Coefficient (xlogp) based Drug Clustering | |
|
|
||
| (3) Hydrogen Bond Donor Count (hbonddonor) based Drug Clustering | (4) Hydrogen Bond Acceptor Count (hbondacc) based Drug Clustering | |
|
|
||
| (5) Rotatable Bond Count (rotbonds) based Drug Clustering | (6) Topological Polar Surface Area (polararea) based Drug Clustering | |
|
|
||
| "RO5" indicates the cutoff set by lipinski's rule of five; "D123AB" colored in GREEN denotes the no violation of any cutoff in lipinski's rule of five; "D123AB" colored in PURPLE refers to the violation of only one cutoff in lipinski's rule of five; "D123AB" colored in BLACK represents the violation of more than one cutoffs in lipinski's rule of five | ||
| Target Poor or Non Binders | Top | |||||
|---|---|---|---|---|---|---|
| Target Poor or Non Binders | ||||||
| Target Regulators | Top | |||||
|---|---|---|---|---|---|---|
| Target-regulating microRNAs | ||||||
| Target-interacting Proteins | ||||||
| Target Profiles in Patients | Top | |||||
|---|---|---|---|---|---|---|
| Target Expression Profile (TEP) | ||||||
| Target-Related Models and Studies | Top | |||||
|---|---|---|---|---|---|---|
| Target Validation | ||||||
| References | Top | |||||
|---|---|---|---|---|---|---|
| REF 1 | Synthesis of bisindolylmaleimide macrocycles, Bioorg. Med. Chem. Lett. 5(18):2093-2096 (1995). | |||||
| REF 2 | Compounds, formulations, and methods of protein kinase C inhibition. US8889672. | |||||
| REF 3 | Substituted pyrido[2,3-d]pyrimidin-7(8H)-ones and therapeutic uses thereof. US8889696. | |||||
| REF 4 | Bisindolylmaleimide inhibitors of protein kinase C. Further conformational restriction of a tertiary amine side chain, Bioorg. Med. Chem. Lett. 4(11):1303-1308 (1994). | |||||
| REF 5 | Multivariate analysis by the minimum spanning tree method of the structural determinants of diphenylethylenes and triphenylacrylonitriles implicate... J Med Chem. 1992 Feb 7;35(3):573-83. | |||||
| REF 6 | Protein kinase C zeta isoform is critical for proliferation in human glioblastoma cell lines. J Neurooncol. 2000 Apr;47(2):109-15. | |||||
| REF 7 | (S)-13-[(dimethylamino)methyl]-10,11,14,15-tetrahydro-4,9:16, 21-dimetheno-1H, 13H-dibenzo[e,k]pyrrolo[3,4-h][1,4,13]oxadiazacyclohexadecene-1,3(2H... J Med Chem. 1996 Jul 5;39(14):2664-71. | |||||
| REF 8 | Inhibitors of protein kinase C. 1. 2,3-Bisarylmaleimides. J Med Chem. 1992 Jan;35(1):177-84. | |||||
| REF 9 | Novel protein kinase C inhibitors: synthesis and PKC inhibition of beta-substituted polythiophene derivatives. Bioorg Med Chem Lett. 1999 Aug 2;9(15):2279-82. | |||||
If You Find Any Error in Data or Bug in Web Service, Please Kindly Report It to Dr. Zhou and Dr. Zhang.

