Target Information
| Target General Information | Top | |||||
|---|---|---|---|---|---|---|
| Target ID |
T08014
(Former ID: TTDI02548)
|
|||||
| Target Name |
TAR DNA binding protein 43 (TARDBP)
|
|||||
| Synonyms |
TDP43; TDP-43; TAR DNA-binding protein 43
Click to Show/Hide
|
|||||
| Gene Name |
TARDBP
|
|||||
| Target Type |
Clinical trial target
|
[1] | ||||
| Disease | [+] 2 Target-related Diseases | + | ||||
| 1 | Acute myeloid leukaemia [ICD-11: 2A60] | |||||
| 2 | Alzheimer disease [ICD-11: 8A20] | |||||
| Function |
RNA-binding protein that is involved in various steps of RNA biogenesis and processing. Preferentially binds, via its two RNA recognition motifs RRM1 and RRM2, to GU-repeats on RNA molecules predominantly localized within long introns and in the 3'UTR of mRNAs. In turn, regulates the splicing of many non-coding and protein-coding RNAs including proteins involved in neuronal survival, as well as mRNAs that encode proteins relevant for neurodegenerative diseases. Plays a role in maintaining mitochondrial homeostasis by regulating the processing of mitochondrial transcripts. Regulates also mRNA stability by recruiting CNOT7/CAF1 deadenylase on mRNA 3'UTR leading to poly(A) tail deadenylation and thus shortening. In response to oxidative insult, associates with stalled ribosomes localized to stress granules (SGs) and contributes to cell survival. Participates also in the normal skeletal muscle formation and regeneration, forming cytoplasmic myo-granules and binding mRNAs that encode sarcomeric proteins. Plays a role in the maintenance of the circadian clock periodicity via stabilization of the CRY1 and CRY2 proteins in a FBXL3-dependent manner.
Click to Show/Hide
|
|||||
| UniProt ID | ||||||
| Sequence |
MSEYIRVTEDENDEPIEIPSEDDGTVLLSTVTAQFPGACGLRYRNPVSQCMRGVRLVEGI
LHAPDAGWGNLVYVVNYPKDNKRKMDETDASSAVKVKRAVQKTSDLIVLGLPWKTTEQDL KEYFSTFGEVLMVQVKKDLKTGHSKGFGFVRFTEYETQVKVMSQRHMIDGRWCDCKLPNS KQSQDEPLRSRKVFVGRCTEDMTEDELREFFSQYGDVMDVFIPKPFRAFAFVTFADDQIA QSLCGEDLIIKGISVHISNAEPKHNSNRQLERSGRFGGNPGGFGNQGGFGNSRGGGAGLG NNQGSNMGGGMNFGAFSINPAMMAAAQAALQSSWGMMGMLASQQNQSGPSGNNQNQGNMQ REPNQAFGSGNNSYSGSNSGAAIGWGSASNAGSGSGFNGGFGSSMDSKSSGWGM Click to Show/Hide
|
|||||
| 3D Structure | Click to Show 3D Structure of This Target | AlphaFold | ||||
| HIT2.0 ID | T25NV9 ; T96QZX | |||||
| Drugs and Modes of Action | Top | |||||
|---|---|---|---|---|---|---|
| Clinical Trial Drug(s) | [+] 2 Clinical Trial Drugs | + | ||||
| 1 | LMT-X | Drug Info | Phase 3 | Acute myeloid leukaemia | [2] | |
| 2 | PP-007 | Drug Info | Phase 1 | Ischemic stroke | [3] | |
| Mode of Action | [+] 1 Modes of Action | + | ||||
| Inhibitor | [+] 1 Inhibitor drugs | + | ||||
| 1 | LMT-X | Drug Info | [1], [4] | |||
| Cell-based Target Expression Variations | Top | |||||
|---|---|---|---|---|---|---|
| Cell-based Target Expression Variations | ||||||
| Different Human System Profiles of Target | Top |
|---|---|
|
Human Similarity Proteins
of target is determined by comparing the sequence similarity of all human proteins with the target based on BLAST. The similarity proteins for a target are defined as the proteins with E-value < 0.005 and outside the protein families of the target.
A target that has fewer human similarity proteins outside its family is commonly regarded to possess a greater capacity to avoid undesired interactions and thus increase the possibility of finding successful drugs
(Brief Bioinform, 21: 649-662, 2020).
Human Tissue Distribution
of target is determined from a proteomics study that quantified more than 12,000 genes across 32 normal human tissues. Tissue Specificity (TS) score was used to define the enrichment of target across tissues.
The distribution of targets among different tissues or organs need to be taken into consideration when assessing the target druggability, as it is generally accepted that the wider the target distribution, the greater the concern over potential adverse effects
(Nat Rev Drug Discov, 20: 64-81, 2021).
Human Pathway Affiliation
of target is determined by the life-essential pathways provided on KEGG database. The target-affiliated pathways were defined based on the following two criteria (a) the pathways of the studied target should be life-essential for both healthy individuals and patients, and (b) the studied target should occupy an upstream position in the pathways and therefore had the ability to regulate biological function.
Targets involved in a fewer pathways have greater likelihood to be successfully developed, while those associated with more human pathways increase the chance of undesirable interferences with other human processes
(Pharmacol Rev, 58: 259-279, 2006).
Biological Network Descriptors
of target is determined based on a human protein-protein interactions (PPI) network consisting of 9,309 proteins and 52,713 PPIs, which were with a high confidence score of ≥ 0.95 collected from STRING database.
The network properties of targets based on protein-protein interactions (PPIs) have been widely adopted for the assessment of target’s druggability. Proteins with high node degree tend to have a high impact on network function through multiple interactions, while proteins with high betweenness centrality are regarded to be central for communication in interaction networks and regulate the flow of signaling information
(Front Pharmacol, 9, 1245, 2018;
Curr Opin Struct Biol. 44:134-142, 2017).
Human Similarity Proteins
Human Tissue Distribution
Human Pathway Affiliation
Biological Network Descriptors
|
|
|
There is no similarity protein (E value < 0.005) for this target
|
|
Note:
If a protein has TS (tissue specficity) scores at least in one tissue >= 2.5, this protein is called tissue-enriched (including tissue-enriched-but-not-specific and tissue-specific). In the plots, the vertical lines are at thresholds 2.5 and 4.
|
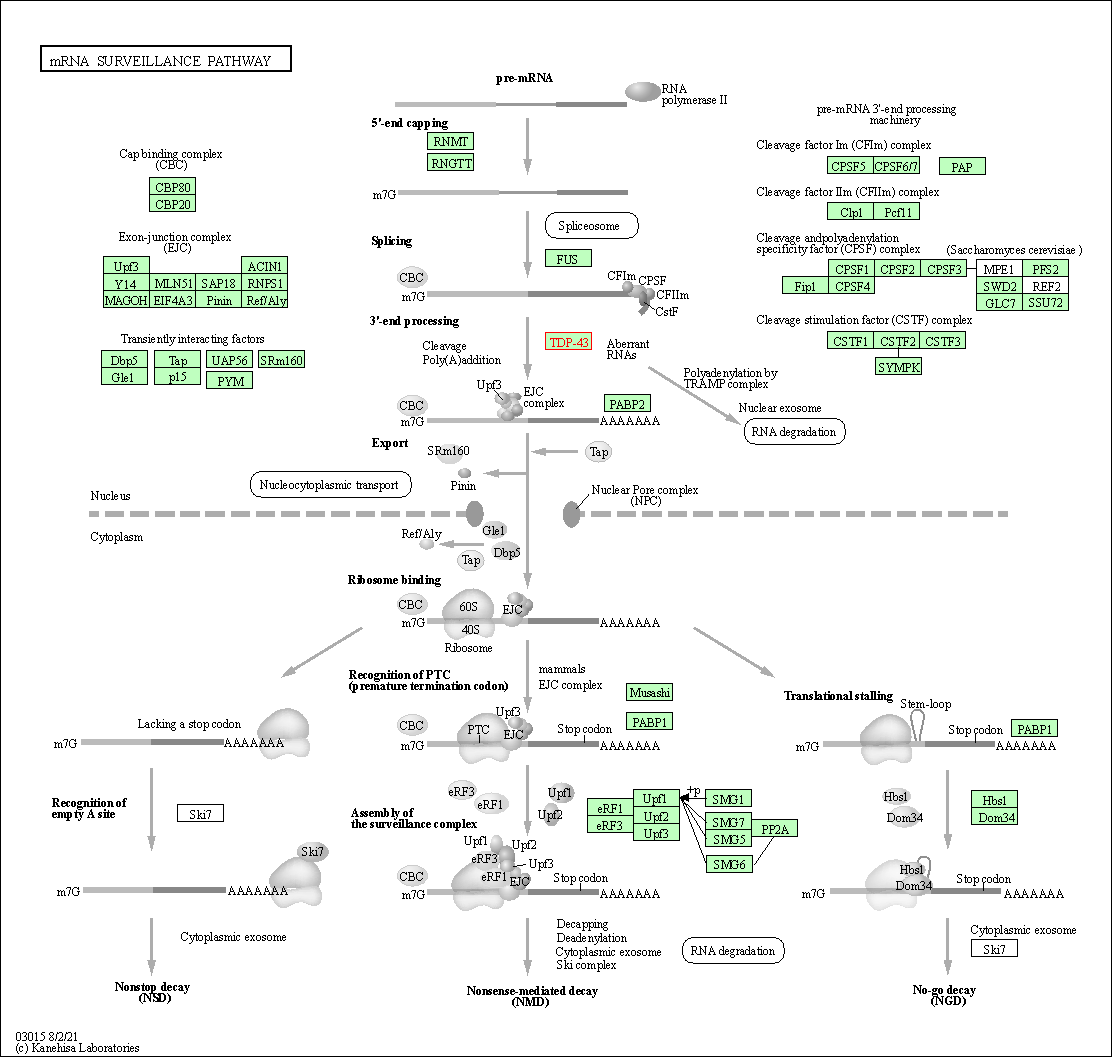
| KEGG Pathway | Pathway ID | Affiliated Target | Pathway Map |
|---|---|---|---|
| mRNA surveillance pathway | hsa03015 | Affiliated Target |

|
| Class: Genetic Information Processing => Translation | Pathway Hierarchy | ||
| Degree | 17 | Degree centrality | 1.83E-03 | Betweenness centrality | 1.66E-04 |
|---|---|---|---|---|---|
| Closeness centrality | 2.14E-01 | Radiality | 1.38E+01 | Clustering coefficient | 6.40E-01 |
| Neighborhood connectivity | 3.22E+01 | Topological coefficient | 2.28E-01 | Eccentricity | 12 |
| Download | Click to Download the Full PPI Network of This Target | ||||
| Target Regulators | Top | |||||
|---|---|---|---|---|---|---|
| Target-regulating microRNAs | ||||||
| Target-interacting Proteins | ||||||
| References | Top | |||||
|---|---|---|---|---|---|---|
| REF 1 | Clinical pipeline report, company report or official report of the Pharmaceutical Research and Manufacturers of America (PhRMA) | |||||
| REF 2 | ClinicalTrials.gov (NCT01689246) Safety and Efficacy Study Evaluating TRx0237 in Subjects With Mild to Moderate Alzheimer's Disease. U.S. National Institutes of Health. | |||||
| REF 3 | ClinicalTrials.gov (NCT04677777) A Randomized, Phase 1, Contemporaneously Controlled, Multicenter Study to Assess the Safety of PP-007 in Subjects With Acute Ischemic Stroke. U.S.National Institutes of Health. | |||||
| REF 4 | Clinical pipeline report, company report or official report of the Pharmaceutical Research and Manufacturers of America (PhRMA) | |||||
If You Find Any Error in Data or Bug in Web Service, Please Kindly Report It to Dr. Zhou and Dr. Zhang.

