Target Information
| Target General Information | Top | |||||
|---|---|---|---|---|---|---|
| Target ID |
T38257
(Former ID: TTDI00078)
|
|||||
| Target Name |
Phosphatase and tensin homolog (PTEN)
|
|||||
| Synonyms |
TEP1; Phosphatidylinositol 3,4,5trisphosphate 3phosphatase and dualspecificity protein phosphatase PTEN; Phosphatidylinositol 3,4,5-trisphosphate 3-phosphatase and dual-specificity protein phosphatase PTEN; Mutated in multiple advanced cancers 1; MMAC1
Click to Show/Hide
|
|||||
| Gene Name |
PTEN
|
|||||
| Target Type |
Literature-reported target
|
[1] | ||||
| Function |
Acts as a dual-specificity protein phosphatase, dephosphorylating tyrosine-, serine- and threonine-phosphorylated proteins. Also acts as a lipid phosphatase, removing the phosphate in the D3 position of the inositol ring from phosphatidylinositol 3,4,5-trisphosphate, phosphatidylinositol 3,4-diphosphate, phosphatidylinositol 3-phosphate and inositol 1,3,4,5-tetrakisphosphate with order of substrate preference in vitro PtdIns(3,4,5)P3 > PtdIns(3,4)P2 > PtdIns3P > Ins(1,3,4,5)P4. The lipid phosphatase activity is critical for its tumor suppressor function. Antagonizes the PI3K-AKT/PKB signaling pathway by dephosphorylating phosphoinositides and thereby modulating cell cycle progression and cell survival. The unphosphorylated form cooperates with AIP1 to suppress AKT1 activation. Dephosphorylates tyrosine-phosphorylated focal adhesion kinase and inhibits cell migration and integrin-mediated cell spreading and focal adhesion formation. Plays a role as a key modulator of the AKT-mTOR signaling pathway controlling the tempo of the process of newborn neurons integration during adult neurogenesis, including correct neuron positioning, dendritic development and synapse formation. May be a negative regulator of insulin signaling and glucose metabolism in adipose tissue. The nuclear monoubiquitinated form possesses greater apoptotic potential, whereas the cytoplasmic nonubiquitinated form induces less tumor suppressive ability. In motile cells, suppresses the formation of lateral pseudopods and thereby promotes cell polarization and directed movement. Tumor suppressor.
Click to Show/Hide
|
|||||
| BioChemical Class |
Phosphoric monoester hydrolase
|
|||||
| UniProt ID | ||||||
| EC Number |
EC 3.1.3.16
|
|||||
| Sequence |
MTAIIKEIVSRNKRRYQEDGFDLDLTYIYPNIIAMGFPAERLEGVYRNNIDDVVRFLDSK
HKNHYKIYNLCAERHYDTAKFNCRVAQYPFEDHNPPQLELIKPFCEDLDQWLSEDDNHVA AIHCKAGKGRTGVMICAYLLHRGKFLKAQEALDFYGEVRTRDKKGVTIPSQRRYVYYYSY LLKNHLDYRPVALLFHKMMFETIPMFSGGTCNPQFVVCQLKVKIYSSNSGPTRREDKFMY FEFPQPLPVCGDIKVEFFHKQNKMLKKDKMFHFWVNTFFIPGPEETSEKVENGSLCDQEI DSICSIERADNDKEYLVLTLTKNDLDKANKDKANRYFSPNFKVKLYFTKTVEEPSNPEAS SSTSVTPDVSDNEPDHYRYSDTTDSDPENEPFDEDQHTQITKV Click to Show/Hide
|
|||||
| 3D Structure | Click to Show 3D Structure of This Target | PDB | ||||
| HIT2.0 ID | T67ZT3 | |||||
| Cell-based Target Expression Variations | Top | |||||
|---|---|---|---|---|---|---|
| Cell-based Target Expression Variations | ||||||
| Different Human System Profiles of Target | Top |
|---|---|
|
Human Similarity Proteins
of target is determined by comparing the sequence similarity of all human proteins with the target based on BLAST. The similarity proteins for a target are defined as the proteins with E-value < 0.005 and outside the protein families of the target.
A target that has fewer human similarity proteins outside its family is commonly regarded to possess a greater capacity to avoid undesired interactions and thus increase the possibility of finding successful drugs
(Brief Bioinform, 21: 649-662, 2020).
Human Tissue Distribution
of target is determined from a proteomics study that quantified more than 12,000 genes across 32 normal human tissues. Tissue Specificity (TS) score was used to define the enrichment of target across tissues.
The distribution of targets among different tissues or organs need to be taken into consideration when assessing the target druggability, as it is generally accepted that the wider the target distribution, the greater the concern over potential adverse effects
(Nat Rev Drug Discov, 20: 64-81, 2021).
Human Pathway Affiliation
of target is determined by the life-essential pathways provided on KEGG database. The target-affiliated pathways were defined based on the following two criteria (a) the pathways of the studied target should be life-essential for both healthy individuals and patients, and (b) the studied target should occupy an upstream position in the pathways and therefore had the ability to regulate biological function.
Targets involved in a fewer pathways have greater likelihood to be successfully developed, while those associated with more human pathways increase the chance of undesirable interferences with other human processes
(Pharmacol Rev, 58: 259-279, 2006).
Biological Network Descriptors
of target is determined based on a human protein-protein interactions (PPI) network consisting of 9,309 proteins and 52,713 PPIs, which were with a high confidence score of ≥ 0.95 collected from STRING database.
The network properties of targets based on protein-protein interactions (PPIs) have been widely adopted for the assessment of target’s druggability. Proteins with high node degree tend to have a high impact on network function through multiple interactions, while proteins with high betweenness centrality are regarded to be central for communication in interaction networks and regulate the flow of signaling information
(Front Pharmacol, 9, 1245, 2018;
Curr Opin Struct Biol. 44:134-142, 2017).
Human Similarity Proteins
Human Tissue Distribution
Human Pathway Affiliation
Biological Network Descriptors
|
|
|
Note:
If a protein has TS (tissue specficity) scores at least in one tissue >= 2.5, this protein is called tissue-enriched (including tissue-enriched-but-not-specific and tissue-specific). In the plots, the vertical lines are at thresholds 2.5 and 4.
|
| KEGG Pathway | Pathway ID | Affiliated Target | Pathway Map |
|---|---|---|---|
| Inositol phosphate metabolism | hsa00562 | Affiliated Target |
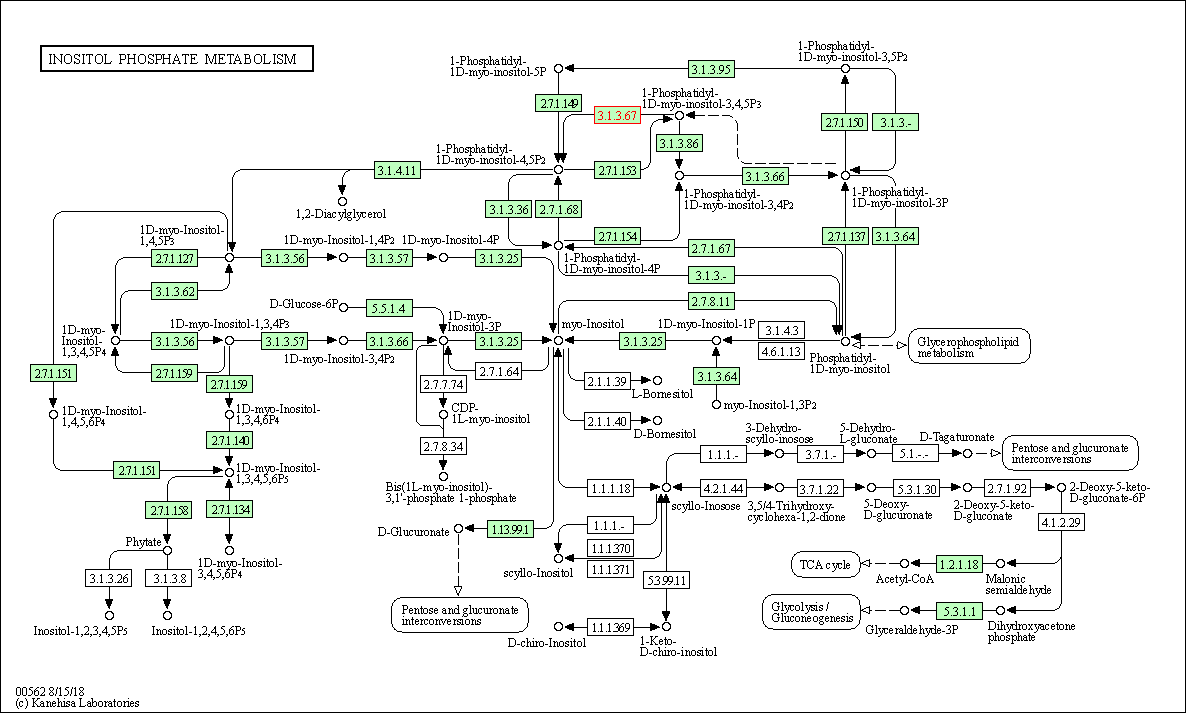
|
| Class: Metabolism => Carbohydrate metabolism | Pathway Hierarchy | ||
| FoxO signaling pathway | hsa04068 | Affiliated Target |
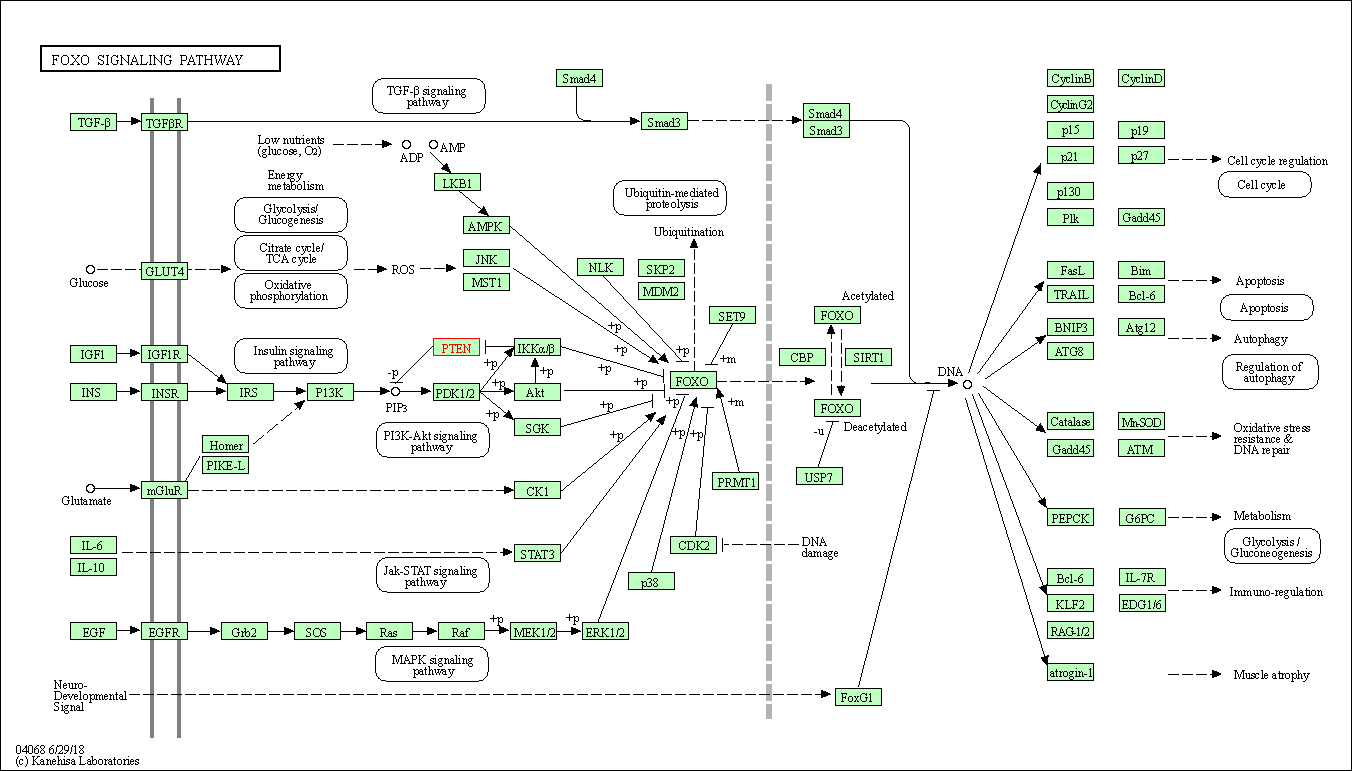
|
| Class: Environmental Information Processing => Signal transduction | Pathway Hierarchy | ||
| Phosphatidylinositol signaling system | hsa04070 | Affiliated Target |
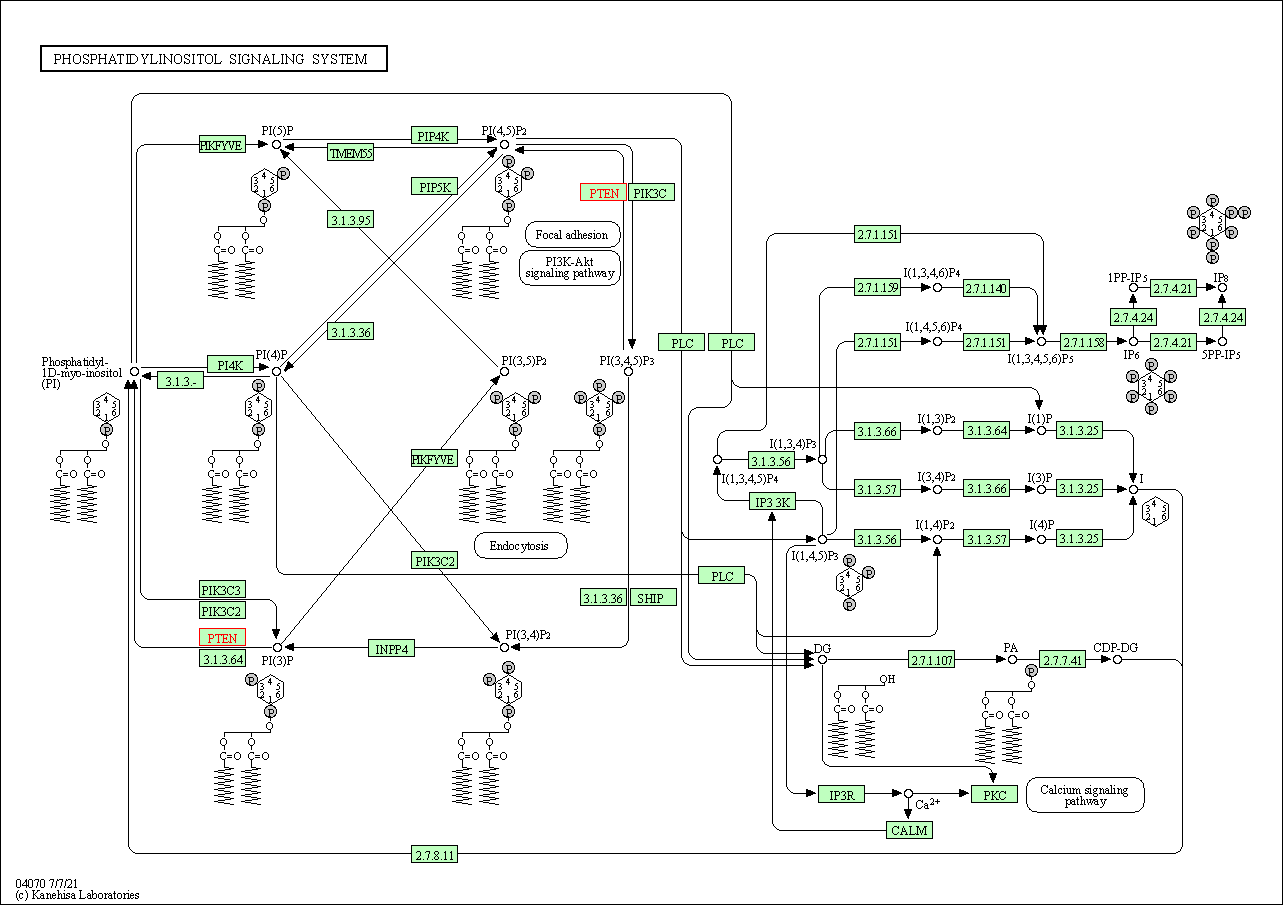
|
| Class: Environmental Information Processing => Signal transduction | Pathway Hierarchy | ||
| Sphingolipid signaling pathway | hsa04071 | Affiliated Target |
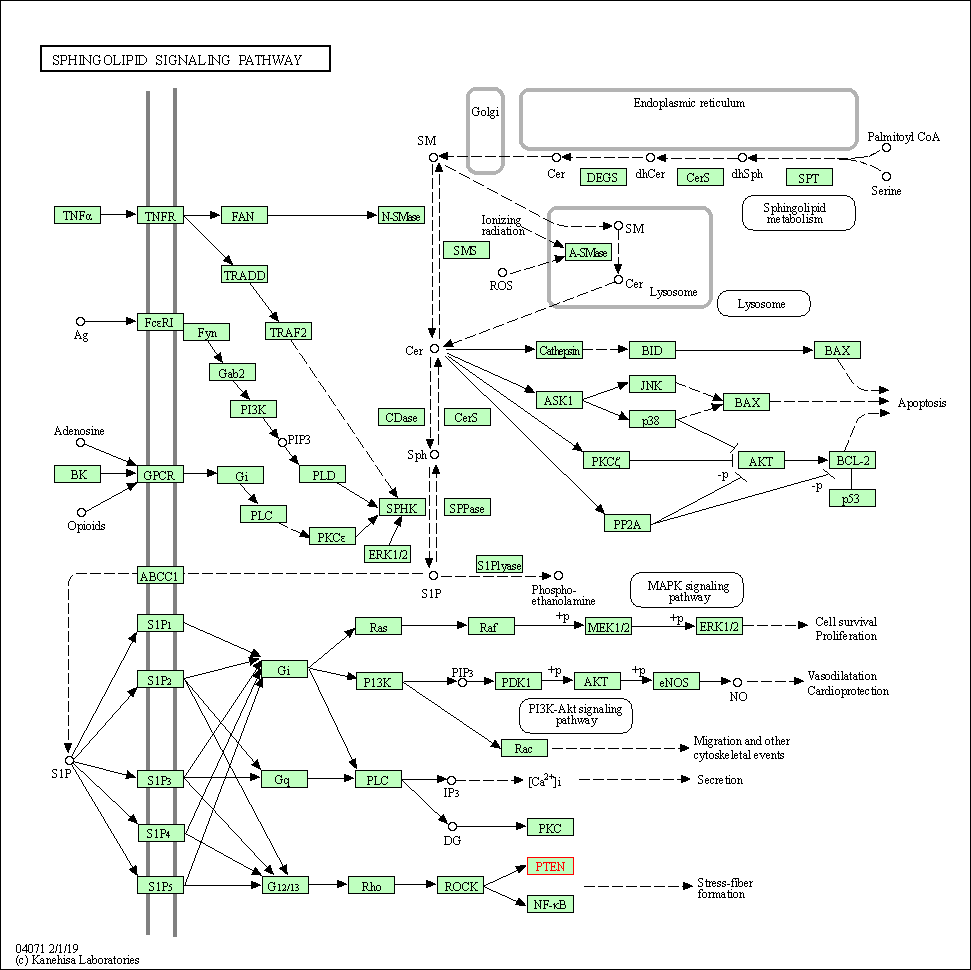
|
| Class: Environmental Information Processing => Signal transduction | Pathway Hierarchy | ||
| p53 signaling pathway | hsa04115 | Affiliated Target |
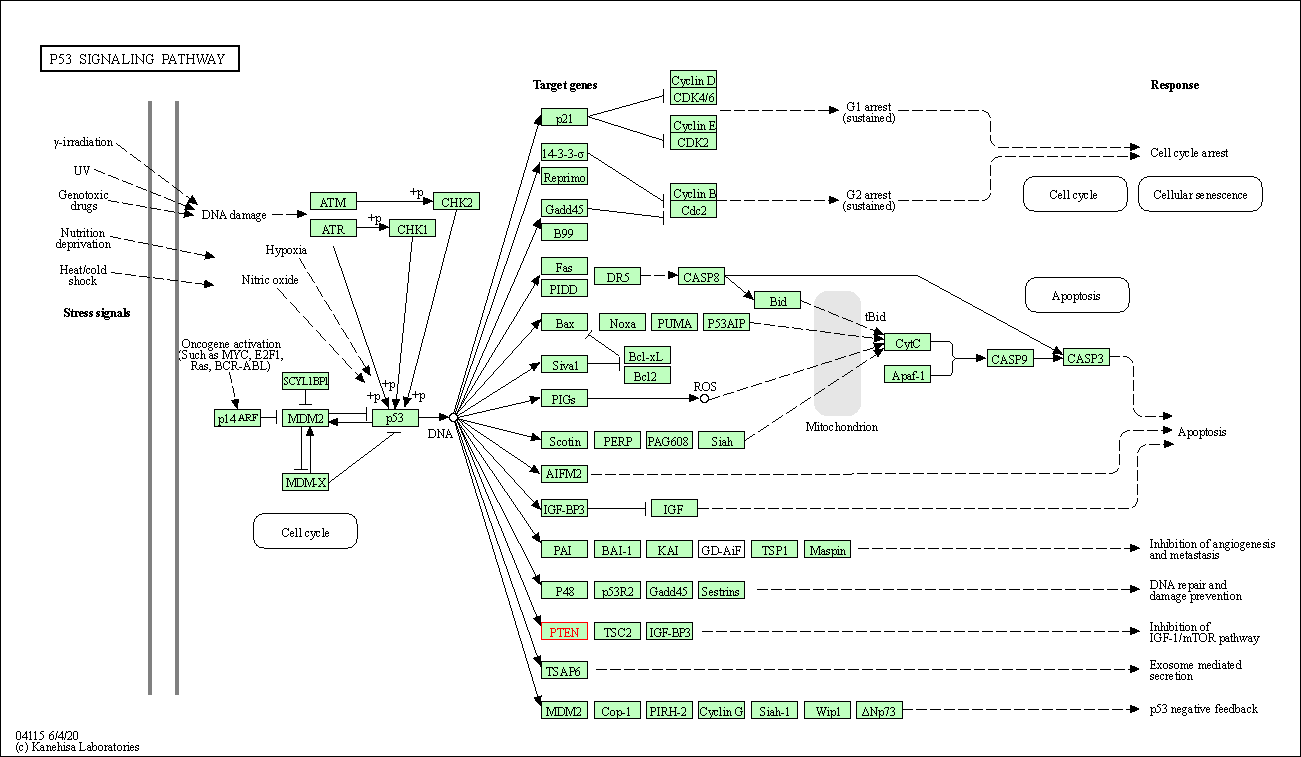
|
| Class: Cellular Processes => Cell growth and death | Pathway Hierarchy | ||
| Autophagy - animal | hsa04140 | Affiliated Target |
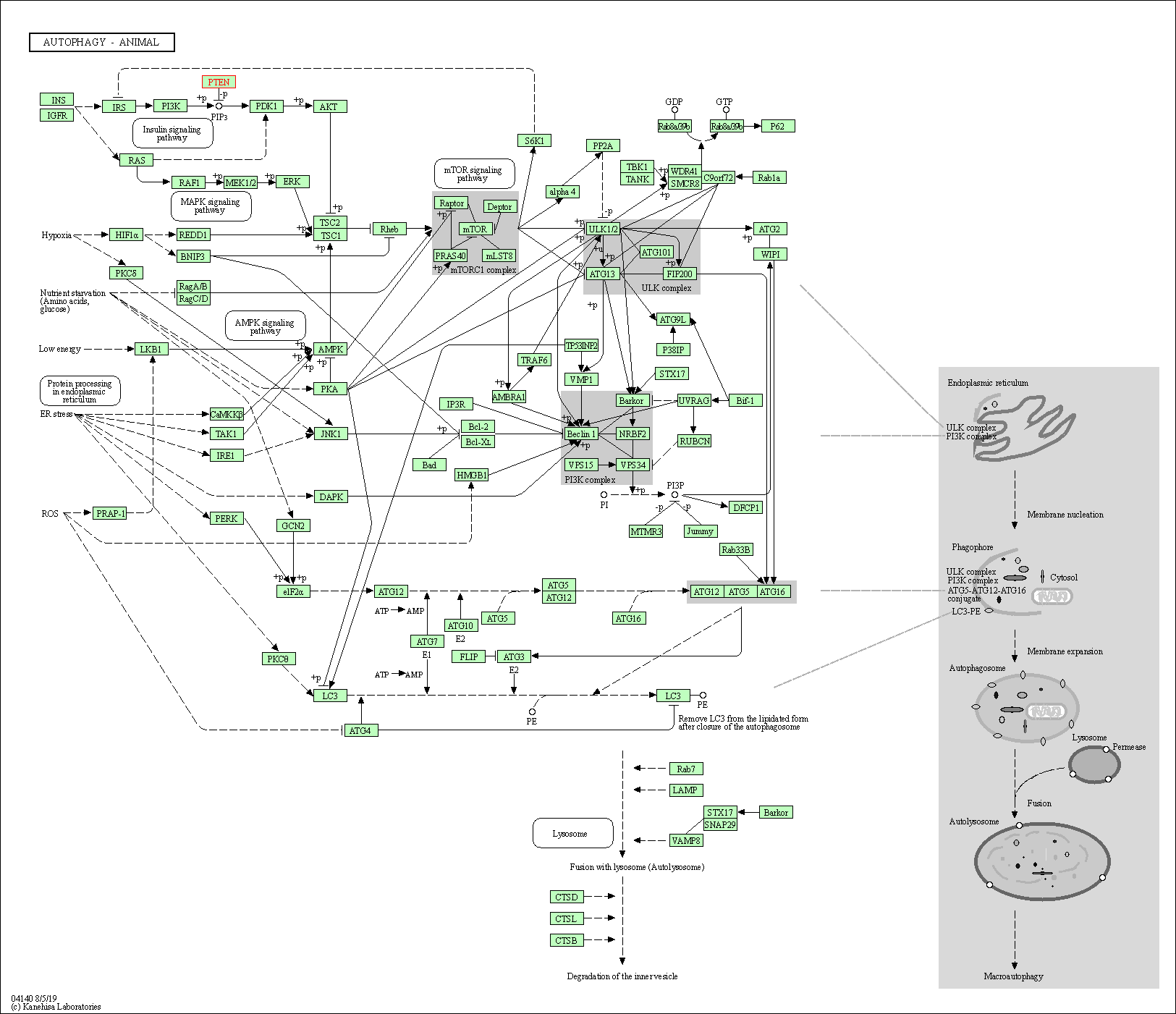
|
| Class: Cellular Processes => Transport and catabolism | Pathway Hierarchy | ||
| mTOR signaling pathway | hsa04150 | Affiliated Target |
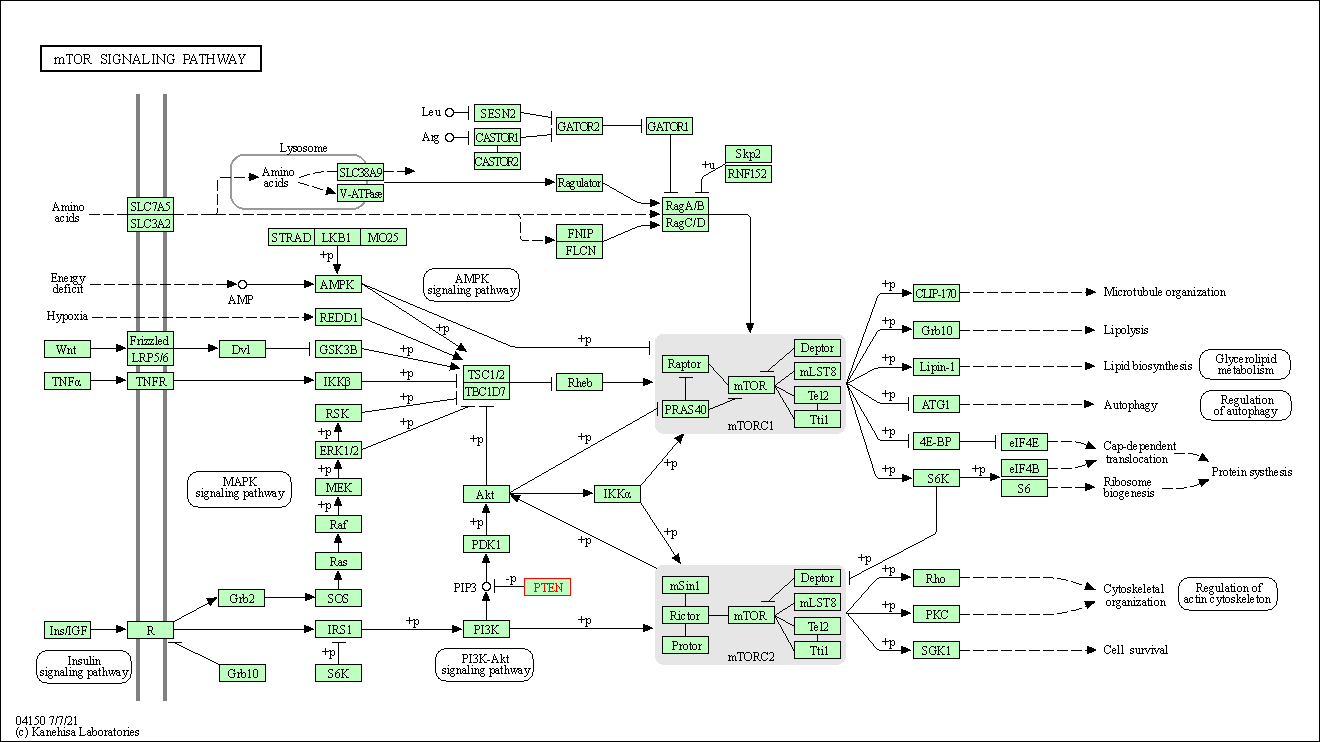
|
| Class: Environmental Information Processing => Signal transduction | Pathway Hierarchy | ||
| PI3K-Akt signaling pathway | hsa04151 | Affiliated Target |
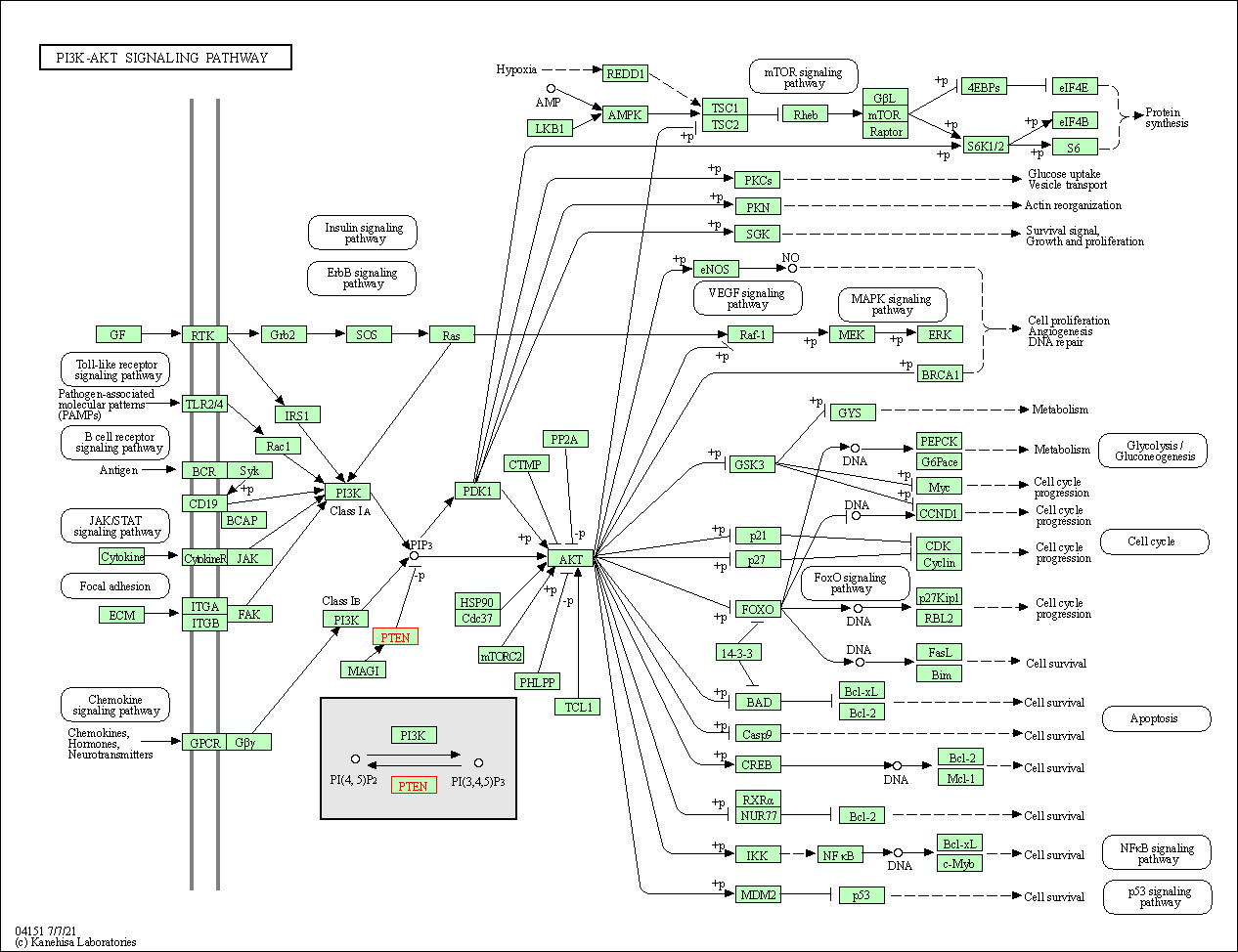
|
| Class: Environmental Information Processing => Signal transduction | Pathway Hierarchy | ||
| Cellular senescence | hsa04218 | Affiliated Target |
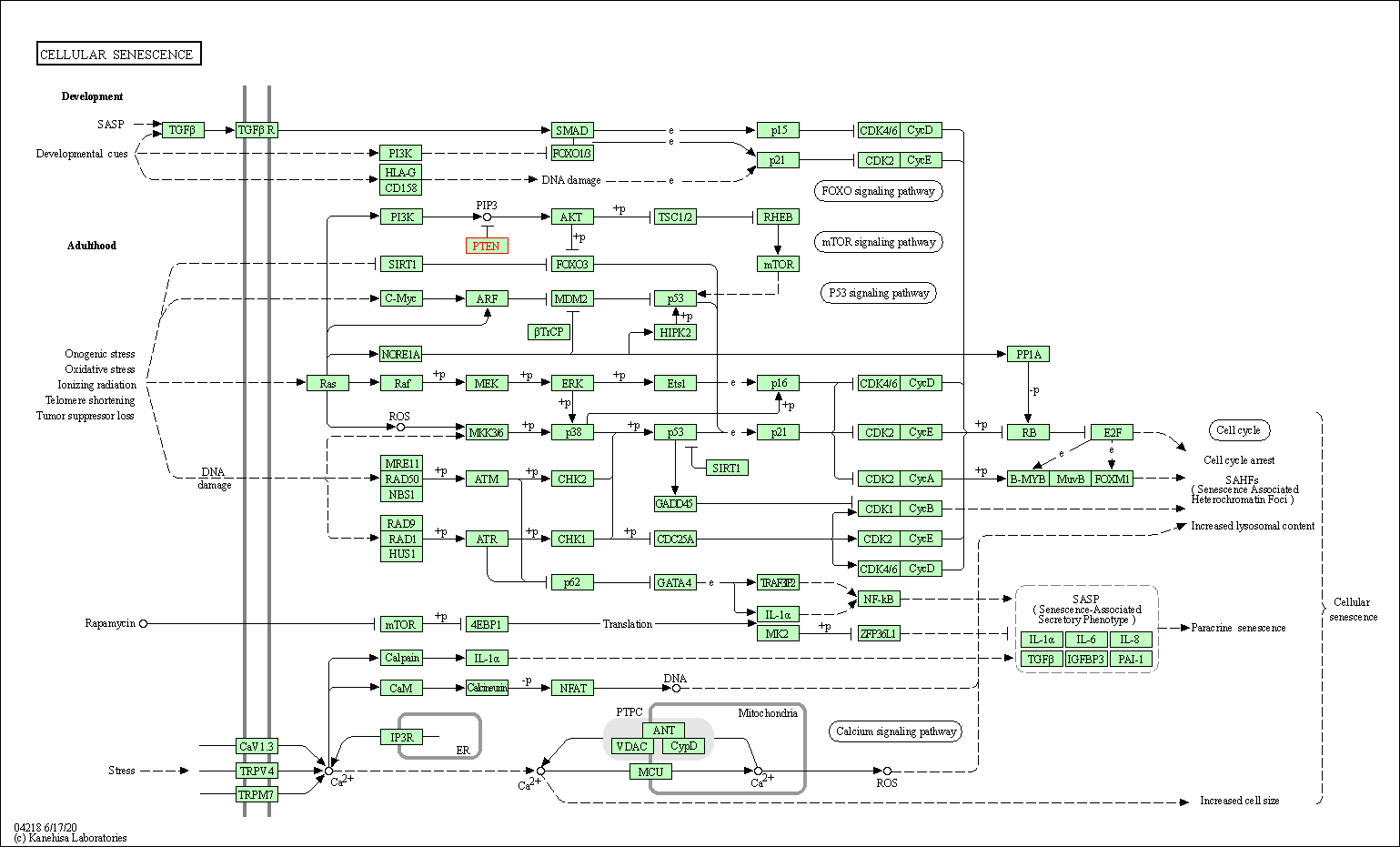
|
| Class: Cellular Processes => Cell growth and death | Pathway Hierarchy | ||
| Focal adhesion | hsa04510 | Affiliated Target |
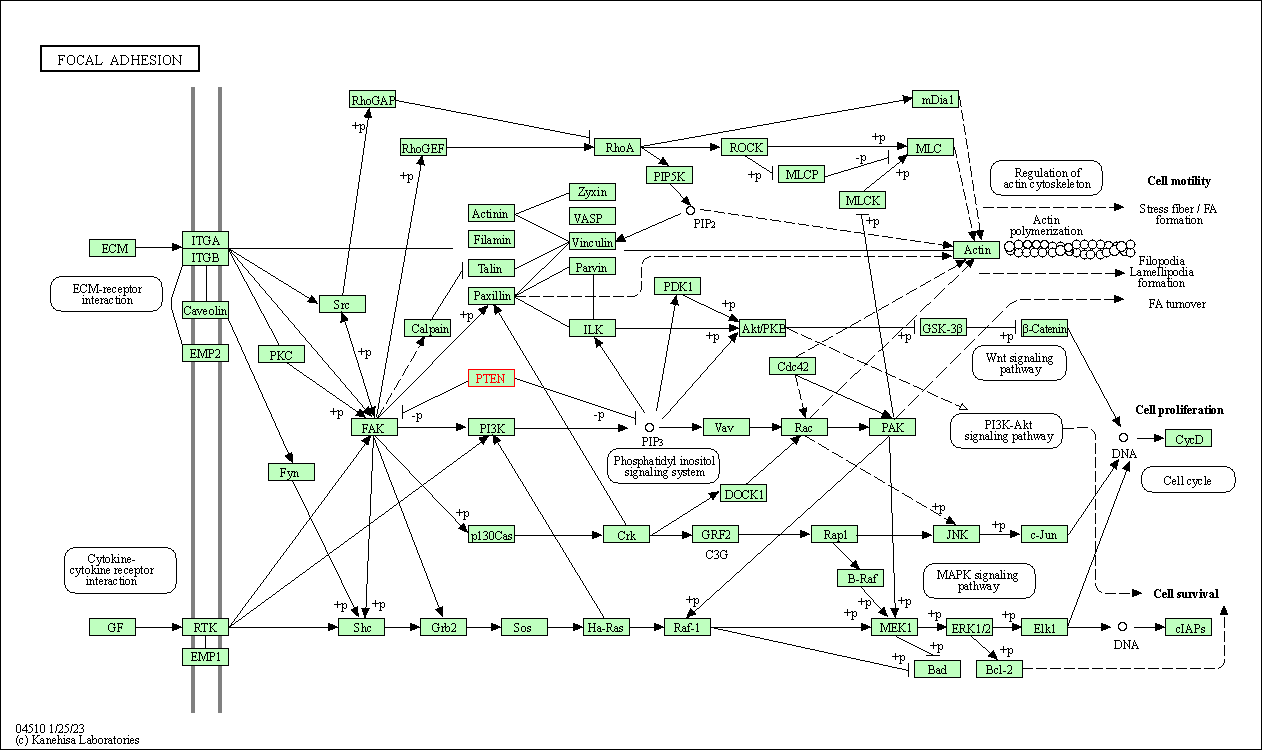
|
| Class: Cellular Processes => Cellular community - eukaryotes | Pathway Hierarchy | ||
| Click to Show/Hide the Information of Affiliated Human Pathways | |||
| Degree | 53 | Degree centrality | 5.69E-03 | Betweenness centrality | 1.02E-02 |
|---|---|---|---|---|---|
| Closeness centrality | 2.71E-01 | Radiality | 1.46E+01 | Clustering coefficient | 5.59E-02 |
| Neighborhood connectivity | 3.79E+01 | Topological coefficient | 3.11E-02 | Eccentricity | 12 |
| Download | Click to Download the Full PPI Network of This Target | ||||
| Target Regulators | Top | |||||
|---|---|---|---|---|---|---|
| Target-regulating microRNAs | ||||||
| Target-regulating Transcription Factors | ||||||
| Target-interacting Proteins | ||||||
| Target Profiles in Patients | Top | |||||
|---|---|---|---|---|---|---|
| Drug Resistance Mutation (DRM) | ||||||
| References | Top | |||||
|---|---|---|---|---|---|---|
| REF 1 | The future therapy of endometrial cancer: microRNA's functionality, capability, and putative clinical application. Arch Gynecol Obstet. 2016 Nov;294(5):889-895. | |||||
If You Find Any Error in Data or Bug in Web Service, Please Kindly Report It to Dr. Zhou and Dr. Zhang.

