Target Information
| Target General Information | Top | |||||
|---|---|---|---|---|---|---|
| Target ID |
T99569
|
|||||
| Target Name |
HUMAN interleukin-10 (IL10)
|
|||||
| Synonyms |
IL-10; Cytokine synthesis inhibitory factor; CSIF
Click to Show/Hide
|
|||||
| Gene Name |
IL10
|
|||||
| Function |
Mechanistically, IL10 binds to its heterotetrameric receptor comprising IL10RA and IL10RB leading to JAK1 and STAT2-mediated phosphorylation of STAT3. In turn, STAT3 translocates to the nucleus where it drives expression of anti-inflammatory mediators. Targets antigen-presenting cells (APCs) such as macrophages and monocytes and inhibits their release of pro-inflammatory cytokines including granulocyte-macrophage colony-stimulating factor /GM-CSF, granulocyte colony-stimulating factor/G-CSF, IL-1 alpha, IL-1 beta, IL-6, IL-8 and TNF-alpha. Interferes also with antigen presentation by reducing the expression of MHC-class II and co-stimulatory molecules, thereby inhibiting their ability to induce T cell activation. In addition, controls the inflammatory response of macrophages by reprogramming essential metabolic pathways including mTOR signaling. Major immune regulatory cytokine that acts on many cells of the immune system where it has profound anti-inflammatory functions, limiting excessive tissue disruption caused by inflammation.
Click to Show/Hide
|
|||||
| BioChemical Class |
Cytokine: interleukin
|
|||||
| UniProt ID | ||||||
| Sequence |
MHSSALLCCLVLLTGVRASPGQGTQSENSCTHFPGNLPNMLRDLRDAFSRVKTFFQMKDQ
LDNLLLKESLLEDFKGYLGCQALSEMIQFYLEEVMPQAENQDPDIKAHVNSLGENLKTLR LRLRRCHRFLPCENKSKAVEQVKNAFNKLQEKGIYKAMSEFDIFINYIEAYMTMKIRN Click to Show/Hide
|
|||||
| 3D Structure | Click to Show 3D Structure of This Target | PDB | ||||
| Cell-based Target Expression Variations | Top | |||||
|---|---|---|---|---|---|---|
| Cell-based Target Expression Variations | ||||||
| Different Human System Profiles of Target | Top |
|---|---|
|
Human Similarity Proteins
of target is determined by comparing the sequence similarity of all human proteins with the target based on BLAST. The similarity proteins for a target are defined as the proteins with E-value < 0.005 and outside the protein families of the target.
A target that has fewer human similarity proteins outside its family is commonly regarded to possess a greater capacity to avoid undesired interactions and thus increase the possibility of finding successful drugs
(Brief Bioinform, 21: 649-662, 2020).
Human Pathway Affiliation
of target is determined by the life-essential pathways provided on KEGG database. The target-affiliated pathways were defined based on the following two criteria (a) the pathways of the studied target should be life-essential for both healthy individuals and patients, and (b) the studied target should occupy an upstream position in the pathways and therefore had the ability to regulate biological function.
Targets involved in a fewer pathways have greater likelihood to be successfully developed, while those associated with more human pathways increase the chance of undesirable interferences with other human processes
(Pharmacol Rev, 58: 259-279, 2006).
Biological Network Descriptors
of target is determined based on a human protein-protein interactions (PPI) network consisting of 9,309 proteins and 52,713 PPIs, which were with a high confidence score of ≥ 0.95 collected from STRING database.
The network properties of targets based on protein-protein interactions (PPIs) have been widely adopted for the assessment of target’s druggability. Proteins with high node degree tend to have a high impact on network function through multiple interactions, while proteins with high betweenness centrality are regarded to be central for communication in interaction networks and regulate the flow of signaling information
(Front Pharmacol, 9, 1245, 2018;
Curr Opin Struct Biol. 44:134-142, 2017).
Human Similarity Proteins
Human Pathway Affiliation
Biological Network Descriptors
|
|
| Protein Name | Pfam ID | Percentage of Identity (%) | E value |
|---|---|---|---|
| Interleukin-22 (IL22) | 28.846 (30/104) | 4.00E-03 |
| KEGG Pathway | Pathway ID | Affiliated Target | Pathway Map |
|---|---|---|---|
| Cytokine-cytokine receptor interaction | hsa04060 | Affiliated Target |
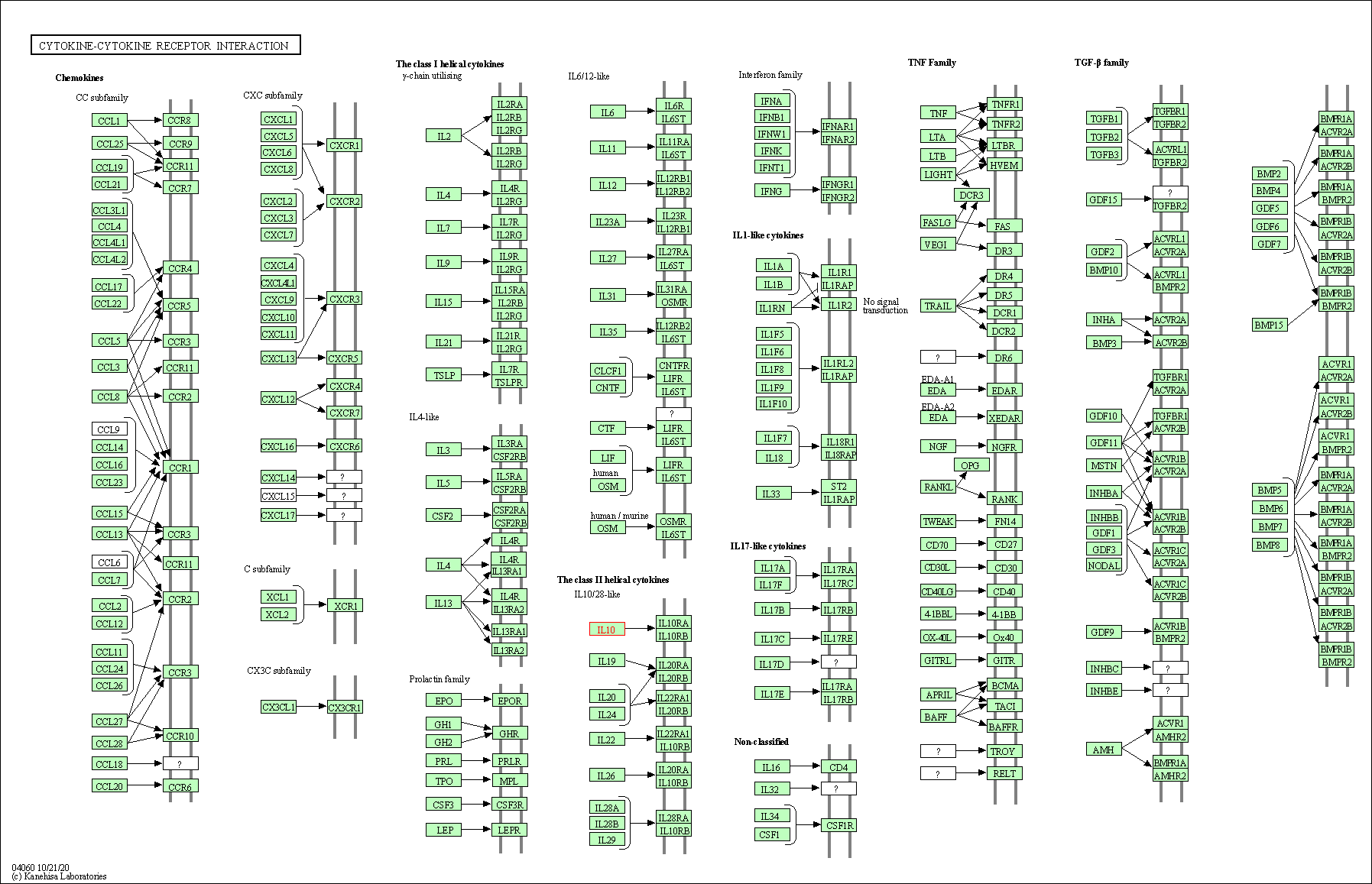
|
| Class: Environmental Information Processing => Signaling molecules and interaction | Pathway Hierarchy | ||
| Viral protein interaction with cytokine and cytokine receptor | hsa04061 | Affiliated Target |
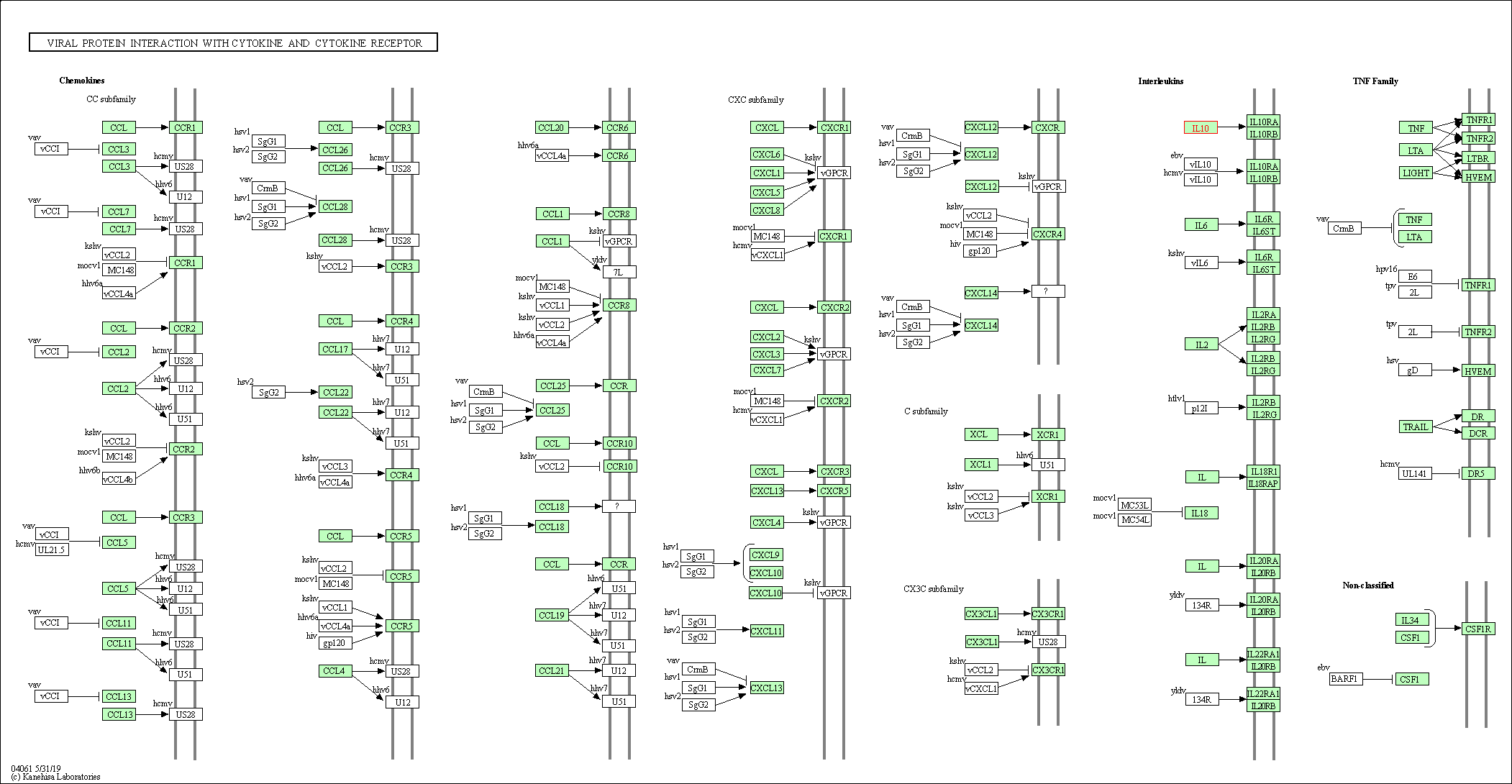
|
| Class: Environmental Information Processing => Signaling molecules and interaction | Pathway Hierarchy | ||
| FoxO signaling pathway | hsa04068 | Affiliated Target |
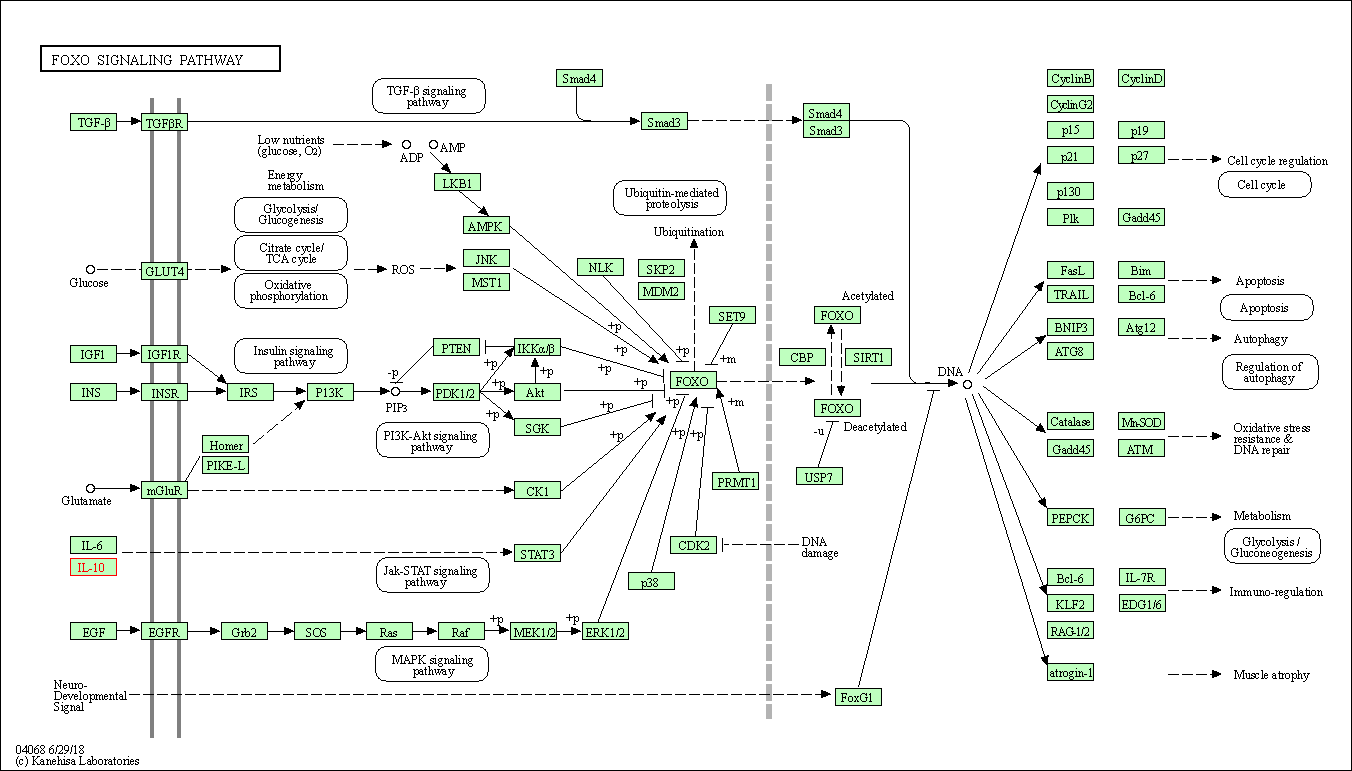
|
| Class: Environmental Information Processing => Signal transduction | Pathway Hierarchy | ||
| C-type lectin receptor signaling pathway | hsa04625 | Affiliated Target |
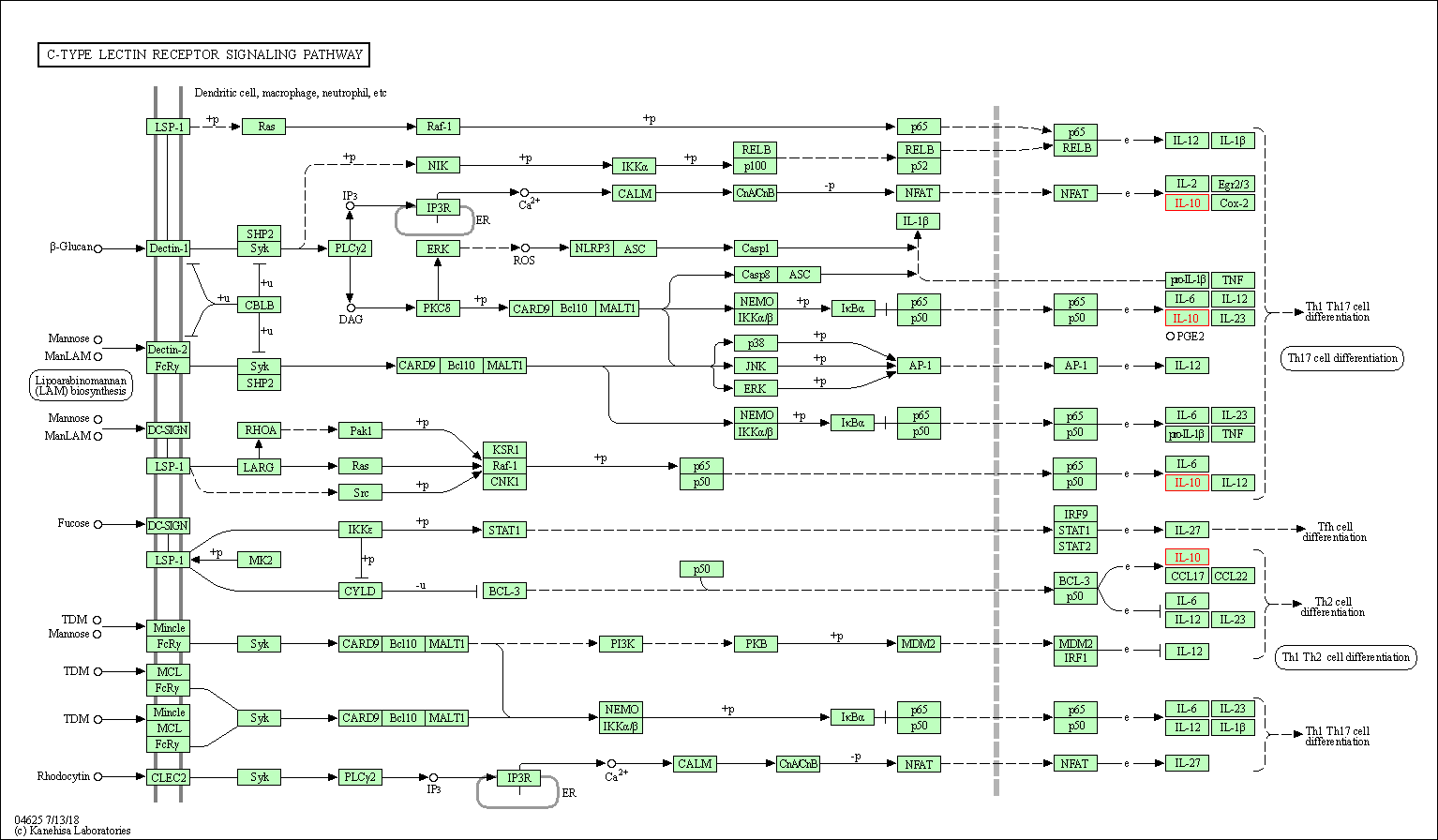
|
| Class: Organismal Systems => Immune system | Pathway Hierarchy | ||
| JAK-STAT signaling pathway | hsa04630 | Affiliated Target |
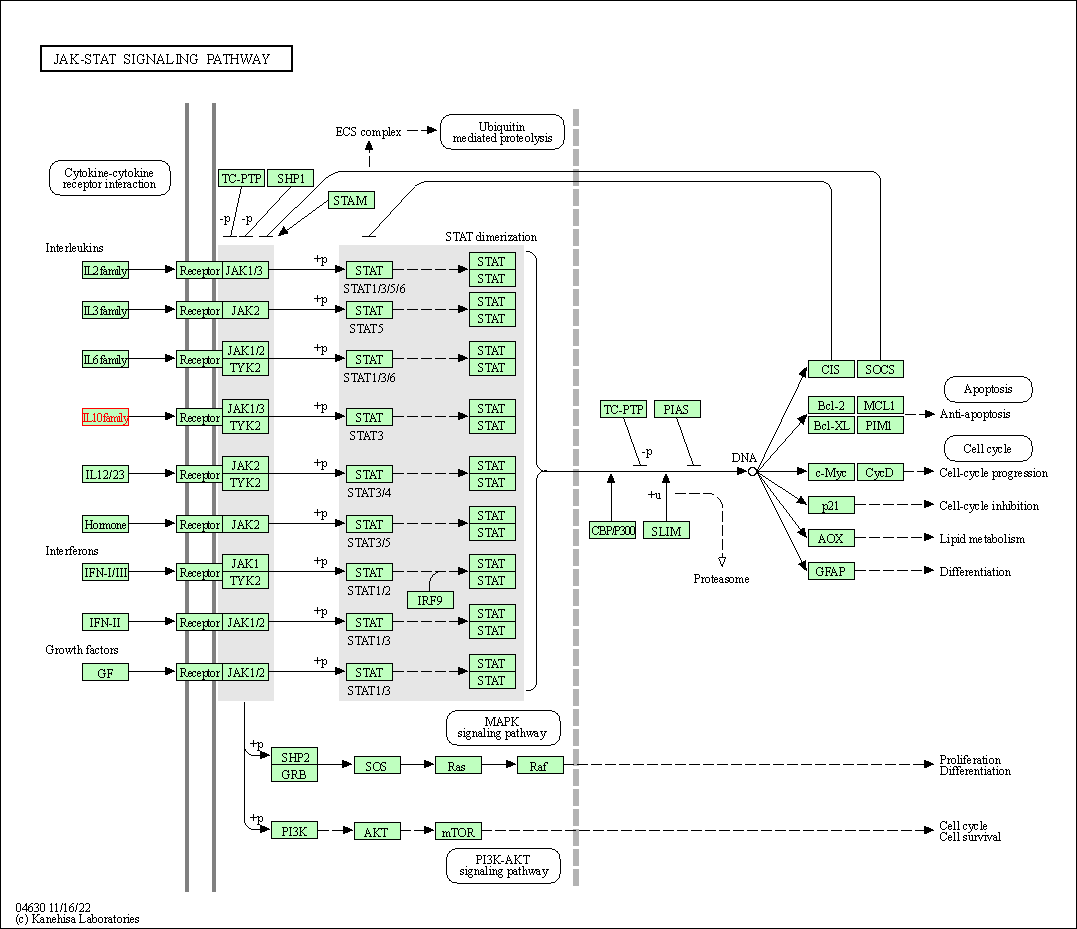
|
| Class: Environmental Information Processing => Signal transduction | Pathway Hierarchy | ||
| T cell receptor signaling pathway | hsa04660 | Affiliated Target |
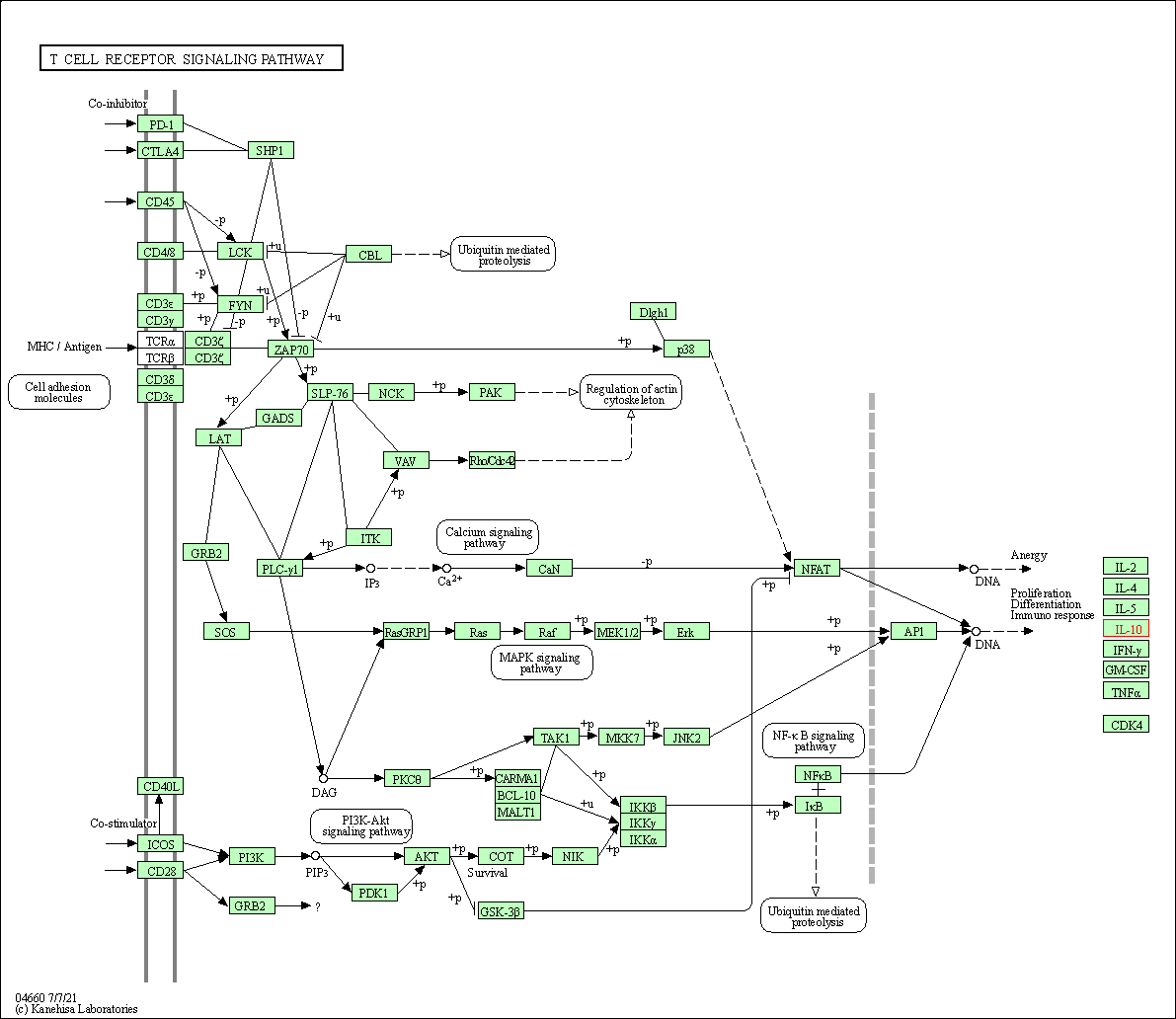
|
| Class: Organismal Systems => Immune system | Pathway Hierarchy | ||
| Intestinal immune network for IgA production | hsa04672 | Affiliated Target |
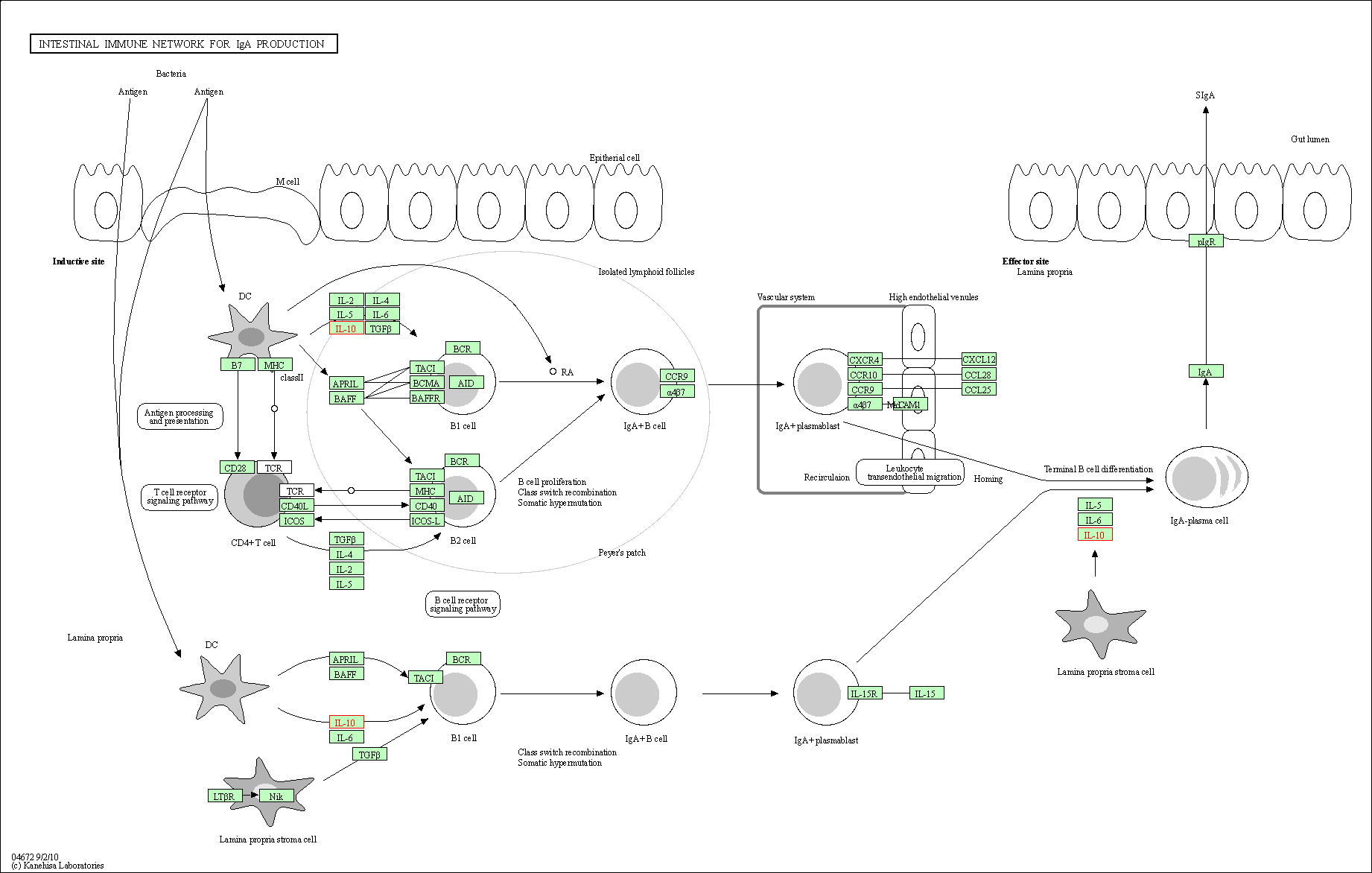
|
| Class: Organismal Systems => Immune system | Pathway Hierarchy | ||
| Click to Show/Hide the Information of Affiliated Human Pathways | |||
| Degree | 55 | Degree centrality | 5.91E-03 | Betweenness centrality | 2.40E-03 |
|---|---|---|---|---|---|
| Closeness centrality | 2.40E-01 | Radiality | 1.42E+01 | Clustering coefficient | 1.70E-01 |
| Neighborhood connectivity | 2.37E+01 | Topological coefficient | 4.86E-02 | Eccentricity | 11 |
| Download | Click to Download the Full PPI Network of This Target | ||||
| References | Top | |||||
|---|---|---|---|---|---|---|
| REF 1 | ClinicalTrials.gov (NCT04363008) COVID-19 Inflammatory Blood Biomarkers for Clinical Management, Prognosis and Evaluation of Interventions | |||||
If You Find Any Error in Data or Bug in Web Service, Please Kindly Report It to Dr. Zhou and Dr. Zhang.

