Target Information
| Target General Information | Top | |||||
|---|---|---|---|---|---|---|
| Target ID |
T77745
(Former ID: TTDI02401)
|
|||||
| Target Name |
Bone marrow stromal antigen 2 (BST2)
|
|||||
| Synonyms |
Tetherin; HM1.24 antigen; CD317; BST2
Click to Show/Hide
|
|||||
| Gene Name |
BST2
|
|||||
| Target Type |
Literature-reported target
|
[1] | ||||
| Function |
IFN-induced antiviral host restriction factor which efficiently blocks the release of diverse mammalian enveloped viruses by directly tethering nascent virions to the membranes of infected cells. Acts as a direct physical tether, holding virions to the cell membrane and linking virions to each other. The tethered virions can be internalized by endocytosis and subsequently degraded or they can remain on the cell surface. In either case, their spread as cell-free virions is restricted. Its target viruses belong to diverse families, including retroviridae: human immunodeficiency virus type 1 (HIV-1), human immunodeficiency virus type 2 (HIV-2), simian immunodeficiency viruses (SIVs), equine infectious anemia virus (EIAV), feline immunodeficiency virus (FIV), prototype foamy virus (PFV), Mason- Pfizer monkey virus (MPMV), human T-cell leukemia virus type 1 (HTLV-1), Rous sarcoma virus (RSV) and murine leukemia virus (MLV), flavivirideae: hepatitis C virus (HCV), filoviridae: ebola virus (EBOV) and marburg virus (MARV), arenaviridae: lassa virus (LASV) and machupo virus (MACV), herpesviridae: kaposis sarcoma- associated herpesvirus (KSHV), rhabdoviridae: vesicular stomatitis virus (VSV), orthomyxoviridae: influenza A virus, and paramyxoviridae: nipah virus. Can inhibit cell surface proteolytic activity of MMP14 causing decreased activation of MMP15 which results in inhibition of cell growth and migration. Can stimulate signaling by LILRA4/ILT7 and consequently provide negative feedback to the production of IFN by plasmacytoid dendritic cells in response to viral infection. Plays a role in the organization of the subapical actin cytoskeleton in polarized epithelial cells. Isoform 1 and isoform 2 are both effective viral restriction factors but have differing antiviral and signaling activities. Isoform 2 is resistant to HIV-1 Vpu-mediated degradation and restricts HIV-1 viral budding in the presence of Vpu. Isoform 1 acts as an activator of NF-kappa-B and this activity is inhibited by isoform 2.
Click to Show/Hide
|
|||||
| BioChemical Class |
Tetherin family
|
|||||
| UniProt ID | ||||||
| Sequence |
MASTSYDYCRVPMEDGDKRCKLLLGIGILVLLIIVILGVPLIIFTIKANSEACRDGLRAV
MECRNVTHLLQQELTEAQKGFQDVEAQAATCNHTVMALMASLDAEKAQGQKKVEELEGEI TTLNHKLQDASAEVERLRRENQVLSVRIADKKYYPSSQDSSSAAAPQLLIVLLGLSALLQ Click to Show/Hide
|
|||||
| 3D Structure | Click to Show 3D Structure of This Target | AlphaFold | ||||
| Cell-based Target Expression Variations | Top | |||||
|---|---|---|---|---|---|---|
| Cell-based Target Expression Variations | ||||||
| Different Human System Profiles of Target | Top |
|---|---|
|
Human Similarity Proteins
of target is determined by comparing the sequence similarity of all human proteins with the target based on BLAST. The similarity proteins for a target are defined as the proteins with E-value < 0.005 and outside the protein families of the target.
A target that has fewer human similarity proteins outside its family is commonly regarded to possess a greater capacity to avoid undesired interactions and thus increase the possibility of finding successful drugs
(Brief Bioinform, 21: 649-662, 2020).
Human Tissue Distribution
of target is determined from a proteomics study that quantified more than 12,000 genes across 32 normal human tissues. Tissue Specificity (TS) score was used to define the enrichment of target across tissues.
The distribution of targets among different tissues or organs need to be taken into consideration when assessing the target druggability, as it is generally accepted that the wider the target distribution, the greater the concern over potential adverse effects
(Nat Rev Drug Discov, 20: 64-81, 2021).
Human Pathway Affiliation
of target is determined by the life-essential pathways provided on KEGG database. The target-affiliated pathways were defined based on the following two criteria (a) the pathways of the studied target should be life-essential for both healthy individuals and patients, and (b) the studied target should occupy an upstream position in the pathways and therefore had the ability to regulate biological function.
Targets involved in a fewer pathways have greater likelihood to be successfully developed, while those associated with more human pathways increase the chance of undesirable interferences with other human processes
(Pharmacol Rev, 58: 259-279, 2006).
Biological Network Descriptors
of target is determined based on a human protein-protein interactions (PPI) network consisting of 9,309 proteins and 52,713 PPIs, which were with a high confidence score of ≥ 0.95 collected from STRING database.
The network properties of targets based on protein-protein interactions (PPIs) have been widely adopted for the assessment of target’s druggability. Proteins with high node degree tend to have a high impact on network function through multiple interactions, while proteins with high betweenness centrality are regarded to be central for communication in interaction networks and regulate the flow of signaling information
(Front Pharmacol, 9, 1245, 2018;
Curr Opin Struct Biol. 44:134-142, 2017).
Human Similarity Proteins
Human Tissue Distribution
Human Pathway Affiliation
Biological Network Descriptors
|
|
|
There is no similarity protein (E value < 0.005) for this target
|
|
Note:
If a protein has TS (tissue specficity) scores at least in one tissue >= 2.5, this protein is called tissue-enriched (including tissue-enriched-but-not-specific and tissue-specific). In the plots, the vertical lines are at thresholds 2.5 and 4.
|
| KEGG Pathway | Pathway ID | Affiliated Target | Pathway Map |
|---|---|---|---|
| Viral life cycle - HIV-1 | hsa03250 | Affiliated Target |
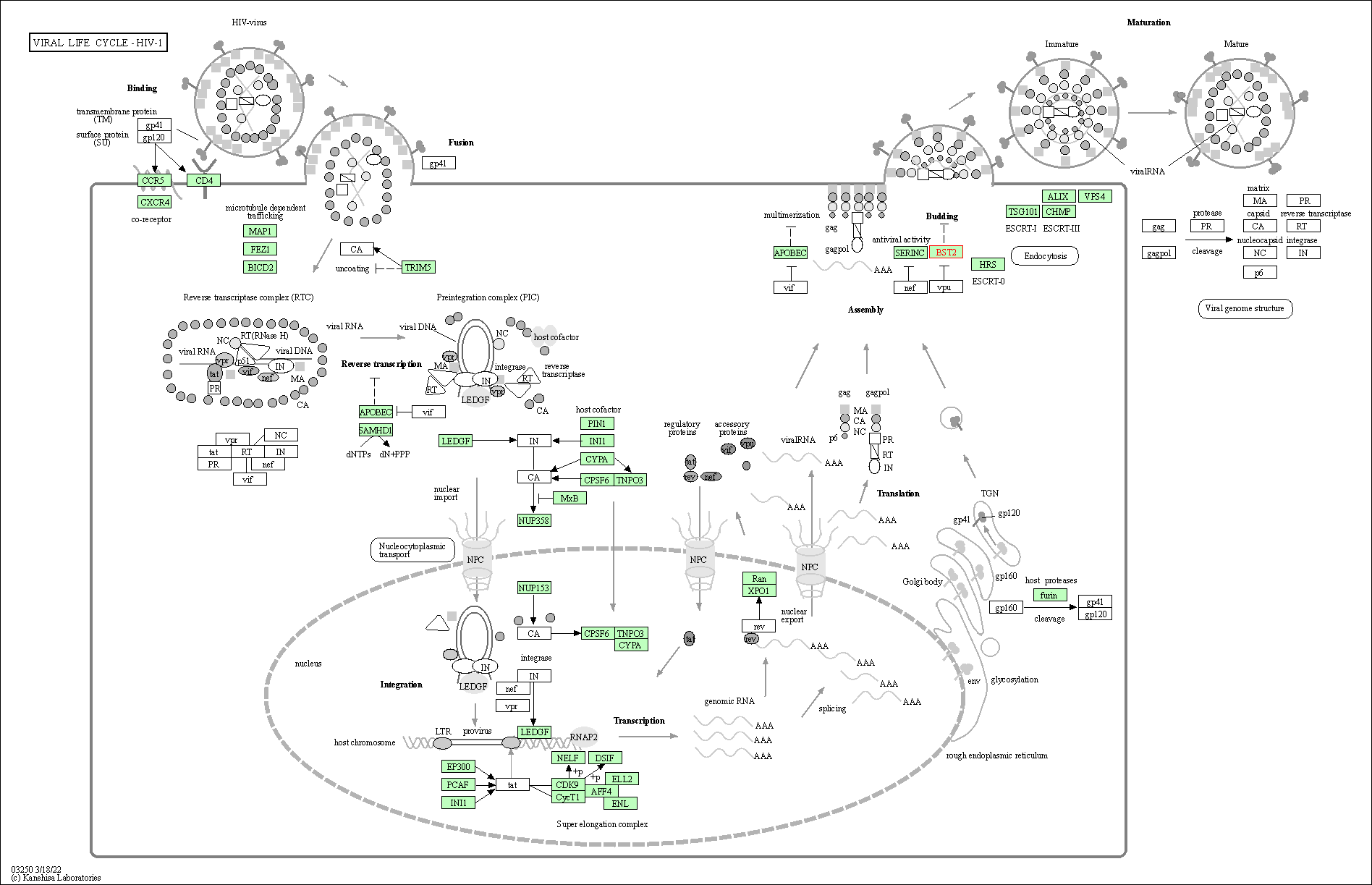
|
| Class: Genetic Information Processing => Information processing in viruses | Pathway Hierarchy | ||
| Degree | 21 | Degree centrality | 2.26E-03 | Betweenness centrality | 9.78E-05 |
|---|---|---|---|---|---|
| Closeness centrality | 2.15E-01 | Radiality | 1.38E+01 | Clustering coefficient | 8.14E-01 |
| Neighborhood connectivity | 3.15E+01 | Topological coefficient | 2.46E-01 | Eccentricity | 12 |
| Download | Click to Download the Full PPI Network of This Target | ||||
| References | Top | |||||
|---|---|---|---|---|---|---|
| REF 1 | Bone marrow stromal antigen 2 expressed in cancer cells promotes mammary tumor growth and metastasis. Breast Cancer Res. 2014 Dec 13;16(6):493. | |||||
If You Find Any Error in Data or Bug in Web Service, Please Kindly Report It to Dr. Zhou and Dr. Zhang.

